SFRP2 Promoter Methylation Analysis in Tumor Tissue, Stool, and Plasma DNA of Patients with Colorectal Cancer
Article Information
Yong-Suk Kim1, Song-Hak Kim2, Myong-Nam Kim3, Un-Rim Sim1*
Department of Molecular Biology, Pyongyang Medical College, Kim Il Sung University, Democratic People’s Republic of Korea
*Corresponding author: Un-Rim Sim, Department of Molecular Biology, Pyongyang Medical College, Kim Il Sung University, Democratic People’s Republic of Korea
Received: 05 September 2019; Accepted: 24 September 2019; Published: 21 October 2019
Citation:
Yong-Suk Kim, Song-Hak Kim, Myong-Nam Kim, Un-Rim Sim. SFRP2 Promoter Methylation Analysis in Tumor Tissue, Stool, and Plasma DNA of Patients with Colorectal Cancer. Archives of Internal Medicine Research 2 (2019): 034-039.
View / Download Pdf Share at FacebookAbstract
Noninvasive method for early detection of human cancers is promising way to provide more quality health service to the population. The serum and stool of colorectal cancer patients often harbor increased free DNA levels, which can potentially be used for cancer detection. Because SFRP2 (secreted frizzled related protein 2) is considered a Wnt inhibitor whose CpGs were frequently hypermethylated in several human cancers, we investigated the frequency of aberrant SFRP2 promoter methylation in primary tumors and serum or stool samples of colorectal cancer patients by MS-PCR. We detected methylation of SFRP2 in 111 of 117 (94.9%) paraffin-embedded colorectal cancer tissues. Fifty-nine of the 68 (86.8%) available serum samples from these cases carried detectable amounts of the methylated SFRP2 promoter. Sixty-one of 68 (89.7%) available stool samples from the same cases carried positive methylation in the promoter region of SFRP2 gene. In contrast, no methylated SFRP2 promoter DNA was detected in serum and stool samples from 60 healthy controls. Only 2 cases of 68 healthy controls showed methylated SFRP2 promoter DNA in their colorectal tissue. But high frequency of methylated SFRP2 gene promoter has no correlation with the clinic-pathologic features. We concluded that SFRP2 methylation analysis appeared to be a noninvasive tumor marker in serum and/or stool DNA
Keywords
SFRP2, Promoter Methylation, Colorectal Cancer
SFRP2 articles SFRP2 Research articles SFRP2 review articles SFRP2 PubMed articles SFRP2 PubMed Central articles SFRP2 2023 articles SFRP2 2024 articles SFRP2 Scopus articles SFRP2 impact factor journals SFRP2 Scopus journals SFRP2 PubMed journals SFRP2 medical journals SFRP2 free journals SFRP2 best journals SFRP2 top journals SFRP2 free medical journals SFRP2 famous journals SFRP2 Google Scholar indexed journals Promoter Methylation articles Promoter Methylation Research articles Promoter Methylation review articles Promoter Methylation PubMed articles Promoter Methylation PubMed Central articles Promoter Methylation 2023 articles Promoter Methylation 2024 articles Promoter Methylation Scopus articles Promoter Methylation impact factor journals Promoter Methylation Scopus journals Promoter Methylation PubMed journals Promoter Methylation medical journals Promoter Methylation free journals Promoter Methylation best journals Promoter Methylation top journals Promoter Methylation free medical journals Promoter Methylation famous journals Promoter Methylation Google Scholar indexed journals Colorectal Cancer articles Colorectal Cancer Research articles Colorectal Cancer review articles Colorectal Cancer PubMed articles Colorectal Cancer PubMed Central articles Colorectal Cancer 2023 articles Colorectal Cancer 2024 articles Colorectal Cancer Scopus articles Colorectal Cancer impact factor journals Colorectal Cancer Scopus journals Colorectal Cancer PubMed journals Colorectal Cancer medical journals Colorectal Cancer free journals Colorectal Cancer best journals Colorectal Cancer top journals Colorectal Cancer free medical journals Colorectal Cancer famous journals Colorectal Cancer Google Scholar indexed journals malignancies articles malignancies Research articles malignancies review articles malignancies PubMed articles malignancies PubMed Central articles malignancies 2023 articles malignancies 2024 articles malignancies Scopus articles malignancies impact factor journals malignancies Scopus journals malignancies PubMed journals malignancies medical journals malignancies free journals malignancies best journals malignancies top journals malignancies free medical journals malignancies famous journals malignancies Google Scholar indexed journals tumor articles tumor Research articles tumor review articles tumor PubMed articles tumor PubMed Central articles tumor 2023 articles tumor 2024 articles tumor Scopus articles tumor impact factor journals tumor Scopus journals tumor PubMed journals tumor medical journals tumor free journals tumor best journals tumor top journals tumor free medical journals tumor famous journals tumor Google Scholar indexed journals serum articles serum Research articles serum review articles serum PubMed articles serum PubMed Central articles serum 2023 articles serum 2024 articles serum Scopus articles serum impact factor journals serum Scopus journals serum PubMed journals serum medical journals serum free journals serum best journals serum top journals serum free medical journals serum famous journals serum Google Scholar indexed journals DNA articles DNA Research articles DNA review articles DNA PubMed articles DNA PubMed Central articles DNA 2023 articles DNA 2024 articles DNA Scopus articles DNA impact factor journals DNA Scopus journals DNA PubMed journals DNA medical journals DNA free journals DNA best journals DNA top journals DNA free medical journals DNA famous journals DNA Google Scholar indexed journals Molecular Biology articles Molecular Biology Research articles Molecular Biology review articles Molecular Biology PubMed articles Molecular Biology PubMed Central articles Molecular Biology 2023 articles Molecular Biology 2024 articles Molecular Biology Scopus articles Molecular Biology impact factor journals Molecular Biology Scopus journals Molecular Biology PubMed journals Molecular Biology medical journals Molecular Biology free journals Molecular Biology best journals Molecular Biology top journals Molecular Biology free medical journals Molecular Biology famous journals Molecular Biology Google Scholar indexed journals histopathological articles histopathological Research articles histopathological review articles histopathological PubMed articles histopathological PubMed Central articles histopathological 2023 articles histopathological 2024 articles histopathological Scopus articles histopathological impact factor journals histopathological Scopus journals histopathological PubMed journals histopathological medical journals histopathological free journals histopathological best journals histopathological top journals histopathological free medical journals histopathological famous journals histopathological Google Scholar indexed journals methylation frequency articles methylation frequency Research articles methylation frequency review articles methylation frequency PubMed articles methylation frequency PubMed Central articles methylation frequency 2023 articles methylation frequency 2024 articles methylation frequency Scopus articles methylation frequency impact factor journals methylation frequency Scopus journals methylation frequency PubMed journals methylation frequency medical journals methylation frequency free journals methylation frequency best journals methylation frequency top journals methylation frequency free medical journals methylation frequency famous journals methylation frequency Google Scholar indexed journals
Article Details
1. Introduction
Colorectal cancer is an important disease with a large morbidity and mortality and increasing health care costs as multi-modality therapy becomes more widespread and new drugs appear. But many of the patients who undergo curative surgical resection will ultimately die of recurrent colorectal cancer, depending mainly on the initial stage of the disease. Thus, earlier detection of colorectal cancer would improve the efficacy of treatment and management of this disease. Therefore, it is vitally important to identify and develop reliable diagnostic markers of early stage colorectal cancer. Several reports show that tumors can release DNA into the circulation or even into the stool, and these DNAs can be often detectable in the serum and stool of cancer patients. The serum of cancer patients contains, on average, 4~40 times more free DNA, compared with normal individual [1].
Aberrant WNT pathway signaling is an early progression event in approximately 90% of CRCs and occurs through mutations mainly of APC and, less often, of CTNNB1 (encoding beta-catenin). These mutations allow ligand independent WNT signaling that culminates in abnormal accumulation of free beta-catenin in the nucleus [2-4]. Secreted frizzled-related proteins (SFRPs) possess a domain similar to one in the WNT-receptor frizzled proteins and can inhibit WNT receptor binding to downregulate pathway signaling during development [5]. Recently, gene silencing of SFRPs, which consist of 5 members (SFRP1-5), by the hypermethylation of their promoter was identified in CRCs and other malignancies [6-8]. Especially, SFRP2 methylation was detected in a high frequency of over 60% of CRCs, including a Lynch syndrome, which suggests that SFRP2 methylation might potentially be useful as a promising sensitive screening marker for the stool-based detection of CRCs and premalignant lesions. We, therefore, investigated macrodissected samples of 117 formalin-fixed and paraffin-embedded colorectal tumor tissues and 68 serum and stool samples from available cases. We also investigated 60 healthy controls in the same way. We proved that serum and stool DNA could be promising marker to detect colorectal cancer in individuals.
2. Materials and Methods
2.1 PatientsTissue specimens from 117 CRCs were obtained from patients with colorectal cancer who had undergone curative surgery at Pyongyang medical school’s hospital, KIM IL SUNG University, Pyongyang, DPRK between 2008 and 2010. Obtained tissue specimens were immediately formalin-fixed and stored as paraffin-embedded until use. Histological examination was performed by the pathology department to determine cancer type and their stage. Among the patients, 68 cases were available to provide serum and stool samples. Sixty healthy volunteers provided their tissue, stool and serum samples.
2.2 DNA preparation and bisulfite modificationDNA was extracted from paraffin-embedded archival materials. DNA concentration was measured using a spectrophotometer (Thermo Fisher Scientific Inc., Wilmington, DE) at 260 nm absorbance. Serum samples were available only from 68 cases and they were collected in tubes with K+EDTA. According to the manufacturer’s instructions, our samples were digested with protease K and DNAs were extracted from 200μl protease K digested samples, following the manufacturer’s protocol. Stool DNA was extracted by phenol-chloroforum-isoamyl alcohol method. Bisulfite modification was carried out using a AxyPrep Multisource Genomic DNA Miniprep Kit.
2.3 Methylation-specific PCR (MS-PCR)The modified DNA was subjected to MS-PCR using specific primers for methylated sequences (forward 5`-GGTCGGAGTTTTTCGGAGTTGCGC -3` and reverse 5`- CCGCTCTCTTCGCTAAATACGACTCG-3`) and unmethylated sequences (forward 5`- TTTTGGGTTGGAGTTTTTTGGAGTTGTGT-3` and reverse 5`-AACCCACTCTCTTCACTAAATACAACTCA-3`), which generate PCR products of 138 and 145 bp, respectively. For PCR, 2 μl of bisulfite-modified DNA was amplified in a total volume of 25 μl containing 1 × PCR buffer (with MgCl2), 12.5 pmol of each primer, 160 μM DNTPs and 0.5 U of Hot-Goldstar Taq-polymerase. PCR conditions were 95°C for 5 min, 35 cycles of 95°C for 1min, annealing with 60°C for 1min and 72°C for 1min followed by a final extension step at 72°C for 10 min. Methylated standard DNA was used as a positive control for methylation, and placenta DNA was used as a negative control. PCR products were electrophoresed on polyacrylamide gels and visualized by silver staining.
2.4 Statistical analysisOne-way analysis, ANOVA with Bonferroni test was used for group comparison. For the statistical analysis, a two-sided value of p<0.05 was considered to indicate statistical significance. Data were analyzed using the SPSS-software 11.5 for windows.
3. Results
3.1 Frequency of SFRP2 promoter methylation in tumor tissue, serum and stoolMethylated SFRP2 promoter was detected in 111 (94.9%) of 117 tumor tissue samples. SFRP2 methylation rate in serum and stool samples are 59 out of 68 (86.8%) and 61 out of 68 (89.7%), respectively. No methylation was detected in serum or stool DNA from 60 healthy controls and only 2 colorectal tissue from healthy controls contained methylated SFRP2 DNA. Detailed information is shown in the Table 1. Figure.1 shows MS-PCR products on polyacrylamide gel electrophoresis.
3.2 Correlation with clinical-pathological factorsIn our group of colorectal carcinomas, there was no association between SFRP2 promoter methylation and age, gender, histopathological type, tumor stage, grade of differentiation. Detailed information is shown in Table 2.
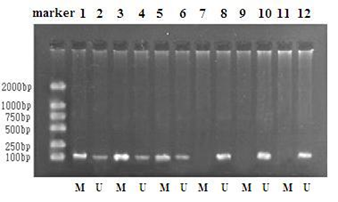
Figure 1: SFRP2 methylation analysis.
|
Sample |
Patients / |
SFRP2 methylated |
SFRP2 unmethylated |
|
Controls |
|||
|
Tissue |
68 |
2 |
66 |
|
117 |
111 |
6 |
|
|
Serum |
60 |
0 |
60 |
|
68 |
59 |
9 |
|
|
Stool |
60 |
0 |
60 |
|
68 |
61 |
7 |
Table 1: SFRP2 promoter methylation frequency.
|
Variable |
N (%) |
SFRP2 methylated |
SFRP2 non-methylated |
|
N (%) |
N (%) |
||
|
Gender |
Male; 56 |
52 (92.9) |
4 (7.1) |
|
Female; 61 |
59 (96.7) |
2 (3.3) |
|
|
Age |
20~40: 19 |
17 (89.5) |
2 (10.5) |
|
41~60: 71 |
69 (97.2) |
2 (2.8) |
|
|
60>: 27 |
25 (92.6) |
2 (7.4) |
|
|
Family history of |
Yes: 16 |
16 (100) |
0 (0) |
|
Cancer patients |
No: 101 |
95 (94.1) |
6 (5.9) |
|
Size of tumor (cm) |
<2.5: 5 |
5 (100) |
0 (0) |
|
2.6~4: 46 |
43 (93.5) |
3 (6.5) |
|
|
4.1>: 66 |
63 (95.5) |
3 (4.5) |
|
|
Tumor stage |
T1~T3a: 60 |
56 (93.3) |
4 (6.7) |
|
T3b~T4: 57 |
55 (96.5) |
2 (3.5) |
|
|
Differentiation |
Low: 6 |
6 (100) |
0 (0) |
|
Medium: 49 |
47 (95.9) |
2 (4.1) |
|
|
High: 62 |
58 (93.5) |
4 (6.5) |
|
|
Histologic type |
Tubular: 72 |
69 (95.8) |
3 (4.2) |
|
Papillary adeno: 31 |
29 (93.5) |
2 (6.5) |
|
|
Signet ring: 14 |
13 (92.8) |
1 (7.2) |
|
|
Invasiveness |
Intramucosal: 2 |
2 (100) |
0 (0) |
|
Submucosa: 19 |
18 (94.7) |
1 (5.4) |
|
|
Tunica: 67 |
64 (95.5) |
3 (4.5) |
|
|
Ambient: 29 |
27 (93.1) |
2 (6.9) |
Table 2: Correlation between clinical-pathological features and methylation rate.
4. Discussion
Our study clearly demonstrates the biological significance of methylation in the region of the SFRP2 promoter in specimens obtained from the colon and rectum. Aberrant methylation in gene promoters has been established as a key mechanism for the inactivation of tumor suppressor genes in human malignancies, including CRCs. Clinically, the identification of genes that are prone to abnormal methylation and that consequently become downregulated is of critical importance, since this is considered to provide a good source of novel biomarkers. The family of SFRP genes, functionally acting as the WNT signaling inhibitors, was recently shown to be a common target of promoter hypermethylation in numerous tumor entities [9-12]. Indeed, we have demonstrated aberrant methylation of the SFRP2 promoter region by analyzing tissue, serum or stool DNA of colorectal cancer patients. This will contribute to the early detection of colorectal cancer in a more quality way.
However, there was no relationship between methylation frequency and clinic-pathological features. Future study must be on the quantification of methylated CpG islands in the promoter region of this gene and clarify the correlation between methylation level and other clinic-pathological features or survival.
References
- Shapiro B, Chakrabarty M, Cohn EM, Leon SA. Determination of circulating DNA levels in patients with benign or malignant gastrointestinal disease. Cancer (Phila) 51 (1983): 2116-2120.
- Fodde R, Smits R, Clevers H. APC, signal transduction and genetic instability in colorectal cancer. Nat Rev Cancer 1 (2001): 55-67.
- Korinek V, Barker N, Morin PJ, van Wichen D, de Weger R, et al. Constitutive transcriptional activation by a beta-catenin-Tcf complex in APC-/- colon carcinoma. Science 275 (1997): 1784-1787.
- Morin PJ, Sparks AB, Korinek V, Barker N, Clevers H, et al. Activation of beta-catenin-Tcf signaling in colon cancer by mutations in beta-catenin or APC. Science 275 (1997): 1787-1790.
- Jones SE, Jomary C. Secreted Frizzled-related proteins:searching for relationships and patterns. Bioessays 24 (2002): 811-820.
- Nagasaka T, Koi M, Kloor M, Gebert J, Vilkin A, et al. Mutations in both KRAS and BRAF may contribute to the meth- ylator phenotype in colon cancer. Gastroenterology 134 (2008): 1950-1960.
- Nishida N, Nishimura T, Nagasaka T, Ikai I, Goel A, et al. Extensive methylation is associated with beta-catenin mutations in hepatocellular carcinoma: evidence for two distinct pathways of human hepatocarcinogenesis. Cancer Res 67 (2007): 4586-4594.
- Veeck J, Noetzel E, Bektas N, Jost E, Hartmann A, et al. Promoter hypermethylation of the SFRP2 gene is a high-frequency alteration and tumor-speci?c epigenetic marker in human breast cancer. Mol Cancer 7 (2008): 83.
- Cheng YY, Yu J, Wong YP, Man EPS, To KF, et al. Frequent epigenetic inactivation of secreted frizzled-related protein 2 (SFRP2) by promoter methylation in human gastric cancer. Br J Cancer 97 (2007): 895-901.
- Esteve P, Bovolenta P. The Advantages and Disadvantages of SFRP1 and SFRP2 Expression in Pathological Events J Biol. Chem 221 (2010): 11-17.
- Jin E-j, Burrusb LW, Ericksona CA. The expression patterns of Wnts and their antagonists during avian eye development. Mech. Dev 116 (2002): 173-176.
- Silva AL, Dawson SN, Arends MJ, Guttula K, Hall N, et al. Boosting Wnt activity during colorectal cancer progression through selective hypermethylation of Wnt signaling antagonists, BMC Cancer 14 (2014): 891.
