Molecular Prediction of SARS-Cov-2 Transmission in Domesticated Livestock
Article Information
Rajarshi Bhattacharya1, Aayatti Mallick Gupta2, Sukhendu Mandal3*, Swadesh R Biswas1*
1Laboratory of Molecular Food Microbiology Department of Botany, Visva-Bharati, Santiniketan-731235, West Bengal, India
2Department of Chemical, Biological & Macro-Molecular Sciences, S. N. Bose National Centre for Basic Sciences, Block-JD, Sector-III, Salt Lake, Kolkata-700106
3Laboratory of Molecular Bacteriology, Department of Microbiology, University of Calcutta, 35, Ballygunge Circular Road, Kolkata–700019, INDIA
*Corresponding author: Sukhendu Mandal, Laboratory of Molecular Bacteriology, Department of Microbiology, University of Calcutta, 35, Ballygunge Circular Road, Kolkata–700019, India.
Swadesh R Biswas, Department of Botany, Visva-Bharati, Santiniketan-731235, West Bengal, India;
Received: 01 December 2021; Accepted: 08 December 2021; Published: 20 December 2021
Citation: Rajarshi Bhattacharya, Aayatti Mallick Gupta, Sukhendu Mandal, Swadesh R Biswas. Molecular Prediction of SARS-Cov-2 Transmission in Domesticated Livestock. Fortune Journal of Health Sciences 4 (2021): 507-519.
View / Download Pdf Share at FacebookAbstract
COVID-19 outbreak caused by severe acute respiratory syndrome Coronavirus-2 (SARS-CoV-2) is an ongoing global pandemic. Although the disease spreads from human to human, the fundamental question concerning the spread of the disease between domesticated animals or from animals to humans remains unanswered. The human angiotensin-converting enzymes 2 (hACE2) receptor, the recognition site for the virus, has orthologs in animals and are structurally and functionally similar to hACE2. This study investigated the nature and strength of interaction between ACE2 of nine pet animals and the receptor binding domain (RBD) of spike protein of SARS-CoV-2. Among nine animals, ACE2 of Oryctolagus and Canis had significantly higher binding affinity of -11.7 Kcal/mol and -13.8 Kcal/mol, burried surface area of 2383 Å2 and 2508 Å2 and Z score of -1.2 and -1.4, respectively. In phylogenetic analysis, the relatedness of ACE2 of Oryctolagus and human was found to be very high. Interestingly, the binding affinity of simulated Oryctolagus-ACE2:RBD was lower in comparison to that of hACE2 as determined from the distance of intermolecular hydrogen-bonds and the energy of interaction. We conclude that like humans, a faster transmission and spread of SARS-CoV-2 among pet animals is not expected to occur.
Keywords
mammalian pets, SARS-CoV-2, COVID-19, Docking, MD simulation
mammalian pets articles, SARS-CoV-2 articles, COVID-19 articles, Docking articles, MD simulation articles, mammalian pets articles mammalian pets Research articles mammalian pets review articles mammalian pets PubMed articles mammalian pets PubMed Central articles mammalian pets 2023 articles mammalian pets 2024 articles mammalian pets Scopus articles mammalian pets impact factor journals mammalian pets Scopus journals mammalian pets PubMed journals mammalian pets medical journals mammalian pets free journals mammalian pets best journals mammalian pets top journals mammalian pets free medical journals mammalian pets famous journals mammalian pets Google Scholar indexed journals SARS-CoV-2 articles SARS-CoV-2 Research articles SARS-CoV-2 review articles SARS-CoV-2 PubMed articles SARS-CoV-2 PubMed Central articles SARS-CoV-2 2023 articles SARS-CoV-2 2024 articles SARS-CoV-2 Scopus articles SARS-CoV-2 impact factor journals SARS-CoV-2 Scopus journals SARS-CoV-2 PubMed journals SARS-CoV-2 medical journals SARS-CoV-2 free journals SARS-CoV-2 best journals SARS-CoV-2 top journals SARS-CoV-2 free medical journals SARS-CoV-2 famous journals SARS-CoV-2 Google Scholar indexed journals COVID-19 articles COVID-19 Research articles COVID-19 review articles COVID-19 PubMed articles COVID-19 PubMed Central articles COVID-19 2023 articles COVID-19 2024 articles COVID-19 Scopus articles COVID-19 impact factor journals COVID-19 Scopus journals COVID-19 PubMed journals COVID-19 medical journals COVID-19 free journals COVID-19 best journals COVID-19 top journals COVID-19 free medical journals COVID-19 famous journals COVID-19 Google Scholar indexed journals Docking articles Docking Research articles Docking review articles Docking PubMed articles Docking PubMed Central articles Docking 2023 articles Docking 2024 articles Docking Scopus articles Docking impact factor journals Docking Scopus journals Docking PubMed journals Docking medical journals Docking free journals Docking best journals Docking top journals Docking free medical journals Docking famous journals Docking Google Scholar indexed journals MD simulation articles MD simulation Research articles MD simulation review articles MD simulation PubMed articles MD simulation PubMed Central articles MD simulation 2023 articles MD simulation 2024 articles MD simulation Scopus articles MD simulation impact factor journals MD simulation Scopus journals MD simulation PubMed journals MD simulation medical journals MD simulation free journals MD simulation best journals MD simulation top journals MD simulation free medical journals MD simulation famous journals MD simulation Google Scholar indexed journals Among nine animals articles Among nine animals Research articles Among nine animals review articles Among nine animals PubMed articles Among nine animals PubMed Central articles Among nine animals 2023 articles Among nine animals 2024 articles Among nine animals Scopus articles Among nine animals impact factor journals Among nine animals Scopus journals Among nine animals PubMed journals Among nine animals medical journals Among nine animals free journals Among nine animals best journals Among nine animals top journals Among nine animals free medical journals Among nine animals famous journals Among nine animals Google Scholar indexed journals BatCoV RaTG13 articles BatCoV RaTG13 Research articles BatCoV RaTG13 review articles BatCoV RaTG13 PubMed articles BatCoV RaTG13 PubMed Central articles BatCoV RaTG13 2023 articles BatCoV RaTG13 2024 articles BatCoV RaTG13 Scopus articles BatCoV RaTG13 impact factor journals BatCoV RaTG13 Scopus journals BatCoV RaTG13 PubMed journals BatCoV RaTG13 medical journals BatCoV RaTG13 free journals BatCoV RaTG13 best journals BatCoV RaTG13 top journals BatCoV RaTG13 free medical journals BatCoV RaTG13 famous journals BatCoV RaTG13 Google Scholar indexed journals ACE2 articles ACE2 Research articles ACE2 review articles ACE2 PubMed articles ACE2 PubMed Central articles ACE2 2023 articles ACE2 2024 articles ACE2 Scopus articles ACE2 impact factor journals ACE2 Scopus journals ACE2 PubMed journals ACE2 medical journals ACE2 free journals ACE2 best journals ACE2 top journals ACE2 free medical journals ACE2 famous journals ACE2 Google Scholar indexed journals
Article Details
1. Introduction
The ongoing COVID-19 pandemic caused by the severe acute respiratory syndrome Coronavirus-2 (SARS-CoV-2) has posed a global challenge for public health with 21.9 Cr infectivity and 45.5 L mortality (https://news.google.com/covid19/) and downfall in the world economy. SARS-CoV-2 belongs to beta coronavirus and known to mainly infect human and few pets as reported [1]. Recent studies showed that SARS-CoV-2 was likely to be originated from bats [2] or pangolins that are predicted to act as natural hosts of SARS-CoV-2[3]. A previous study showed that SARS-CoV-2 replicates poorly in dogs and pigs but cats remain as permissive to the infection [4]. The genome of SARS-CoV-2 shares about 80% identities with that of SARS-CoV and is about 96% identical to the bat coronavirus, BatCoV RaTG13 [5]. A further threat is thus expected to come in post pandemic era as from unknown pet host the pandemic situation can reinitiate at anytime from anywhere. We are addressing this question to reveal the upcoming sources of threat of SARS-CoV-2 by performing in-silico structural studies.
The structure of the trimeric spike protein (S) of SARS-CoV-2, the major factor for host cell infection has been determined [6-8]. It is cleaved by host proteases into the S1 subunit, which contains the receptor-binding domain (RBD), and S2, which mediates fusion of the virion with cellular membranes [9]. Proteolytic cleavage of S at two sites is required to execute its fusion activity. Primarily cleavage is performed by the serine trans membrane protease TMPRSS2, but in its absence the lysosomal cathepsin protease L can also does the same [10].
The primary physiological role of ACE2 is in the maturation of angiotensin, a peptide hormone that stimulates vasoconstriction and blood pressure. ACE2 is a type I membrane protein expressed in lungs, heart, kidneys, and intestine [11]. Down regulation of ACE2 is related to cardiovascular diseases [12]. Total length of ACE2 consists of an N-terminal peptidase domain and a C-terminal collectrin-like domain that ends with a single trans-membrane helix and a ~40-residue intra cellular segment [13, 14]. The peptidase domain (PD) of ACE2 cleaves angiotensine1 to produce angiotensine-(1-9), which is then processed by other enzymes to become angiotensine-(1-7). ACE2 can also directly process angiotensine 2 to give angiotensine-(1-7) [15].
The neck domain and minor interface of peptidase domain are involved in the formation of ACE2 dimer. A monomer of RBD interacts with PD of ACE2 mainly through polar interactions. Arch-shaped a1 helix of the ACE2-PD is interacted by an extended loop region of the RBD [7]. Residue glutamine 493 in SARS-CoV-2 is compatible with hotspot lysine 31 of human ACE2. Second, aspragine 501 of RBD can interact with lysine 353 of human ACE2 [16]. Active residues of hACE2 are 31K, 35E, 38D, 82M and 353K which interact with L455, F486, Q493, S494 and N501 residues of RBD [16].
Many reports on SARS-CoV-2 described the biochemical nature of interaction between human ACE2 and RBD. However, such approach of molecular interaction between animal ACE2 and RBD predicting the transmission of SARS-CoV-2 during the pandemic and in the post-pandemic situation in the domesticated animals is required. In the present study, we focused on eliciting binding interaction of ACE2 receptor from farm animals like Bos, Capra, Ovis, Camalus, Mus and Sus, pet animals, Felis, Canis and Oryctolagus with the receptor binding protein (RBD) of SARS-CoV-2 through molecular docking followed by MD simulation of Oryctolagus ACE2::RBD complex. The information obtained from this study might be useful to understand the possibility of pet-animals to be intermediate hosts of SARS-CoV-2.
2. Materials and Method
2.1 Data mining
Full length amino acids sequence of ACE2 of Homo sapience [accession no.: NP_001358344.1] and few mammalian representatives belonging in same socio-economically-niche of human i.e. Felis catus [accession no.: NP_001034545.1], Camalus ferus [accession no.:XP_006194263.1], Bos taurus [accession no.: XP_005228486.1], Capra hircus [accession no.:NP_001277036.1], Ovis aries [accession no.: XP_011961657.1], Canis lupus familiaris [accession no.:NP_001158732.1], Sus scrofa [accession no.:XP_020935033.1:1-770], Oryctolagus cuniculus [accession no.: XP_002719891.1:12-805], Mus musculus [accession no.: NP_001123985.1] were retrieve from National Center for Biotechnology Information protein database [https://www.ncbi.nlm.nih.gov/protein/].
2.2. Alignment and phylogenetic analysis of ACE2
Multiple Sequence alignment (MSA) of all of the sequences has been performed using the ClustalW of clustal omega web server of the Euoropean Bioin-formatics Institute (EMBL-EBI) [17]. Esprit 3 software[18] was used to represent the MSA using BLOSUM 62 algorithm. The phylogenetic analysis was accomplished through MSA using the neighbor joining algorithm in the MEGA-X (version 10.0.5) [19]. The neighbor-joining phylogenies were estim-ated by using MSA, and the number of bootstraps was 1000. The Poisson correction model and gamma-distributed pattern were used.
2.3. Homology modeling of ACE2
Homology modeling of ACE2 of all retrieved pet sequences were built by using the SWISS-MODEL [20] Web Server, human ACE2 [SMTL ID: 6m18.1] selected as template. The steriochemical property of the built model was evaluated by Ramachandran plot using Volume, Area, Dihedral Angle Reporter (VADAR) server [21]. The structural superim-position of all ACE2 models was performed to visua-lize the 3-D structural differences at domain label by using read scoring matrix in PyMOL software [22].
2.4. Molecular Docking
Molecular Docking was performed to test SARS-CoV-2 spike binding affinity among pet animals to investigate probable pet host next to human. The solvated docking software, HADDOCK [23], was used in this study to dock between the solvated RBD domain of spike protein of SARS-CoV-2 and retrieved mammals ACE2. Molecular docking was performed in HADDOCK server to generate water refined model. Most reliable model was selected by lowest HADDOCK score value. The score is calculated as
HADDOCKscore =1.0 * Evdw + 0.2 * Eelec + 1.0 * Edesol +0.1 *Eair
Where Evdw is the intermolecular van der waals energy, Eelec the intermolecular electrostatic energy, Edesol represents an empirical desolvation energy. The easy interface was utilized since no restrains are defined. Critical residues of ACE2 and RBD domain were used to dock in the HADDOCK web server. In both proteins, the residues surrounding the active residues were selected as passive in HADDOCK. Active residues are the amino acid residues from the two interacting proteins binding sites that take part in direct interaction with other protein partner while passive residues are the residues that can interact indirectly in HADDOCK. Two main types of interactions are established upon docking: H-bonding and hydrophobic interactions. In analogy with any spontaneous process, protein-protein binding occurs only when the change in Gibbs free energy (ΔG) of the system is negative when the system reaches an equilibrium state at constant pressure and temp-erature. Because the protein-protein association extent is determined by the magnitude of the negative ΔG, it can be considered that ΔG determines the stability of any given protein-protein complex. Here we had used Prodigy@Bonvin lab web server for calculating ΔG of our docking study at 25°C and others in default run. PRODIGY software[24] was used to predict the binding affinity of pet mammals. Grand Average of Hydropathy score of ACE2 was calculated by using Expasy Protparam web server. A hydropathy scale has been composed wherein the hydrophilic and hydrophobic properties of each of the 20 amino acid side-chains is taken into consideration. The more positive the value, the more hydrophobic are the amino acids located in that region of the protein.
2.5. Comparative dynamic propensity
The predicted model of Oryctolagus ACE2-CoV2 complex was used as the starting models to check their dynamic behaviour in explicit water model. The spike protein RBD domain is composed of 194 residues, while the ACE2 protein contains 580 residues from the N terminal domain. The simulation parameterization and equilibration were prepared for complex structure using GROMACS version 2018.1. Each system was solvated in SPC216 (simple point charge) water molecule [25] and sodium chloride ion were added to neutralize the system. Approximately, each system was composed of about 255152 atoms (human) and 238363 (Oryctolagus) that were parametrized with the GROMOS96 53a6 force-field [26] After energy minimization using the steepest descent algorithm, each system was equilibrated at temperature 300 K, which was maintained by v-rescale (modified Berendsen thermostat) scheme with 2 fs coupling constant in the NVT ensemble (constant volume and temperature) for 100.0 ps under periodic boundary conditions with harmonic restraint forces applied to the complex molecules 100 kJ mol-1 nm-2. In the subsequent step, the harmonic restraints were removed and the NPT ensembles were simulated at one atmosphere pressure (105 Pa) and 300 k. The pressure was maintained by isotropic Berendsen, with a compressibility of 4.5e-5 bar-1 and coupling time constant of 100 ps. The simulation trajectories were propagated to 1 ns using the GROMACS 2018.1 package.
3. Results
3.1. Alignment and phylogenetic analysis of ACE2
Felis ACE2 protein sequence is the closest to the human ACE2, with 85.22% identity. In contrast ACE2 of Bos, Capra, Ovis, Camalus, Sus, Mus, Oryctolagus, Canis share 81%, 81.72%, 81.72%, 83.23%, 82.34%, 85.14%, 83.46% with human ACE2 (Table S1).
Active residues of human ACE2 involved in binding with RBD of SARS-CoV-2 are 31K, 35E, 38D, 82M and 353K. The animals, Bos, Capra, Ovis and Sus, each with 82T in respect to human 82M and the rest of the active residues are remaining the same as human. Felis and Camalus are having 38E, 82T and 31E, 82T respectively in respect to active residues of hACE2 in the corresponding positions. Canis is with same active residues like Felis. Oryctolagus and Mus are with 82T, 353H and 31N, 82S respectively in ACE2 (Fig. S1 and Table 1)
Table 1: Comparative active residues of human and others pet are tabulated with position and hydrophobicity index
|
Name |
Residue position of interacting hACE2 |
||||||||||||||
|
31 residue HI |
35 residue HI |
38 residue HI |
82 residue HI |
353 residue HI |
similarity with hACE2 |
||||||||||
|
Human |
K |
-0.98 |
E |
-1.4 |
D |
-1.12 |
M |
0.07 |
K |
-1.16 |
5/5 |
||||
|
Felis |
K |
-0.98 |
E |
-1.4 |
E |
-1.5 |
T |
-0.02 |
K |
-1.16 |
4/5 |
||||
|
Camalus |
E |
-0.94 |
E |
-1.3 |
D |
-1.5 |
T |
-0.8 |
K |
-0.56 |
3/5 |
||||
|
Bos |
K |
-0.98 |
E |
--1.4 |
D |
-1.5 |
T |
-0.8 |
K |
-0.56 |
4/5 |
||||
|
Capra |
K |
-0.9 |
E |
-1.4 |
D |
-1.5 |
T |
-0.8 |
K |
-0.56 |
4/5 |
||||
|
Ovis |
K |
-0.9 |
E |
-1.4 |
D |
-1.5 |
T |
-0.8 |
K |
-0.56 |
4/5 |
||||
|
Canis |
K |
-0.5 |
E |
-0.8 |
E |
-1.2 |
T |
-0.5 |
K |
-0.5 |
3/5 |
||||
|
Sus |
K |
-0.21 |
E |
-0.6 |
D |
-0.4 |
T |
-0.6 |
K |
-1.16 |
4/5 |
||||
|
Oryctolagus |
K |
0.14 |
E |
-0.37 |
D |
-0.95 |
T |
-0.86 |
H |
-0.20 |
3/5 |
||||
|
Mus |
N |
0.97 |
E |
-1.4 |
D |
-1.5 |
S |
-0.53 |
K |
-1.89 |
3/5 |
||||
HI stands for hydrophobicity index
Phylogenetic tree was built by using MEGA-X software. Neighbor joining phylogenetic tree was found to consist of two nodes. One node is with human, Oryctolagus, Mus and another one is with Felis, Camalus, Bos, Capra, Ovis, Canis, Sus. So human and rodents family members are belonging to the same node (Fig. 1). This study is based on primary sequence similarity. Whether phylogentically close group does show binding efficiency same or not was addressed by homology modeling followed by docking and MD simulations. The mode of transmission from human to pet can be predicted from the present study.
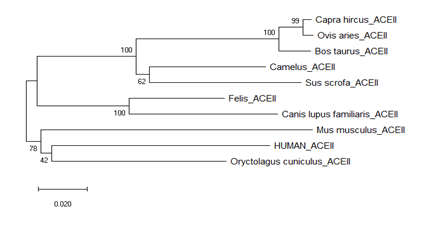
Figure 1:Phylogenetic analysis of sequence of ACE2 of pet animals and human. The ACE2 sequence of human, Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus, Mus were analyzed.
3.2. Homology modeling of ACE2
Homology model were built by using SWISS-MODEL Web Server. Steriochemical properties of built ACE2 model was validated by using Ramachandran plot. Homology model of human, Mus, Sus, Camalus have none of the amino acid residues in the disallowed region. Felis, Canis and Oryctolagus consist of only one disallowed amino acid. The ACE2 structures from rest of the animals have two disallowed amino acids. All models generated by using same template of human ACE2 [SMTL ID: 6m18.1] were superimposed which indicated C-alpha RMSD 0.191 Å in PyMOL (Fig. S2). Structurally, all ACE2 models were turned out to be more or less same. The binding affinity at residue level was then estimated by docking of the models.
Table 2: Comparative affinity of interaction between ACE2 and RBD of SARS-CoV-2
|
Name |
Binding Affinity ΔG(Kcal/mol) |
GRAVY |
Burried Surface area (Å2) |
Z score |
|
Human |
-11.0 |
-0.375 |
2092.4 |
-1.5 |
|
Felis |
-11.9 |
-0.436 |
2383.3 |
-1.2 |
|
Camalus |
-10.8 |
-0.370 |
1872.5 |
-1.4 |
|
Bos |
-11.0 |
-0.445 |
1863.8 |
-1.0 |
|
Capra |
-10.4 |
-0.462 |
1872.5 |
-1.4 |
|
Ovis |
-10.4 |
-0.465 |
2303.7 |
-1.0 |
|
Canis |
-13.8 |
-0.43 |
2508.8 |
-1.4 |
|
Sus |
-10.7 |
-0.323 |
2028.5 |
-1.1 |
|
Oryctolagus |
-11.7 |
-0.444 |
2383.3 |
-1.2 |
|
Mus |
-10.9 |
-0.416 |
1869.1 |
-1.3 |
3.3 Molecular Docking
In our docking study, RBD domain of SARS-CoV-2 was fixed for all pet ACE2 docking run. So calculated ΔG of ACE2 of human, Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus, Mus complexed with RBD is -11.0 Kcal/mol, -11.9Kcal/mol, -10.8 Kcal/mol, -11.0 Kcal/mol, -10.4 kcal/mol,-10.4Kcal/mol, -13.8 Kcal/mol, -10.7 Kcal/mol, -11.7 kcal/mol, -10.9 kcal/mol respectively.
GRAVY score of human, Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus and Mus was calculated as -0.375, -0.0436, -0.370, -0.445, -0.462, -0.465, -0.43, -0.323, -0.444, -0.416 respectively.
Best HADDOCK model was compared within retrieved ACE2 sequences of pets by three parameters of Z-score, Buried surface area, Cluster size. The Z-score indicates how many standard deviations from the average of the cluster is located in terms of score (the more negative the better score). Z-score of human, Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus and Mus was predicted as -1.5,-1.2,-1.4,-1.0,-1.4,-1.0,-1.4,-1.1,-1.2,-1.3. Buried surface area and cluster size partially refers the binding score (more large magnitude is better). Buried surface area of human, Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus and Mus was calculated as 2092.4, 2383.3, 1872.5, 1863.8, 1872.5, 2303.7, 2508.8, 2028.5, 2383.3, 1869.1 (Å2). Cluster size of human, Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus and Mus was also predicted as 58, 187, 78, 258, 60, 14, 360, 18, 343, 18 (Å2).
Based on our molecular interaction study, ACE2 of Canis, Felis and Oryctolagus was found to show higher binding affinity than hACE2 with RBD.
3.4. Simulation of Oryctolagus ACE2 with RBD
Molecular dynamics simulation was carried out to investigate the binding affinity of pet representative Oryctolagus. Simulation of hACE2::RBD was also performed on a same platform to avoid any confusion during comparative analysis. After molecular dynamics simulation of Oryctolagus, only Lys31 and Glu35 of Oryctolagus ACE2 (oACE2) were established hydrogen bonds. Human ACE2::RBD complex was established by formation of stable bonds like Lys31::Tyr453, Glu35::Gln493, Asp38::Thr500, Met82::Phe490 with 2.9 Å, 1.7 Å, 1.9 Å, 2.7 Å, 5.0 Å respectively (Table 3) (Fig. 2).
Table 3: Intermolecular interactions between ACE2 (human and Oryctolagus) and RBD spike glycoprotein of SARS-Cov-2. ACE2 residues are marked in bold
|
Complex |
Interacting residues |
Distance |
Bond type |
|
hACE2::RBD |
LYS31::TYR453 |
2.93135 |
Hydrogen Bond |
|
GLU35::GLN493 |
1.72328 |
Hydrogen Bond |
|
|
GLU35::TYR505 |
1.72907 |
Hydrogen Bond |
|
|
GLU35::TYR495 |
4.82212 |
Electrostatic |
|
|
ASP38::TYR495 |
1.75994 |
Hydrogen Bond |
|
|
ASP38::THR500 |
1.9246 |
Hydrogen Bond |
|
|
ASP38::TYR505 |
2.71356 |
Hydrogen Bond |
|
|
MET82::PHE486 |
2.68056 |
Hydrogen Bond |
|
|
MET82::PHE490 |
5.01727 |
Pi-sulphur |
|
|
MET82::PHE486 |
5.44892 |
Hydrophobic |
|
|
oACE2::RBD |
LYS31::SER443 |
2.06218 |
Hydrogen Bond |
|
LYS31::VAL445 |
5.04797 |
Hydrophobic |
|
|
GLU35::LYS444 |
3.32549 |
Electrostatic |
|
|
GLU35::VAL445 |
2.26932 |
Hydrogen Bond |
|
|
GLU35::GLY446 |
2.19039 |
Hydrogen Bond |
|
|
GLU35::GLY447 |
1.91034 |
Hydrogen Bond |
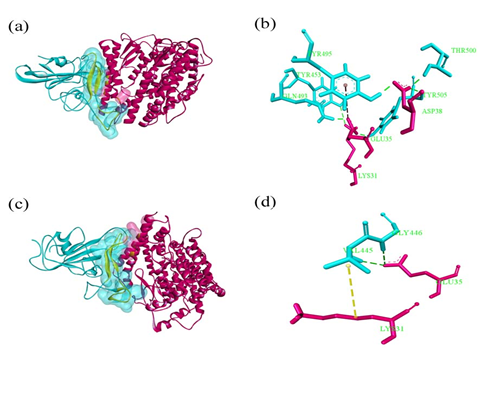
Figure 2: Simulated ensemble of (a)-(b) hACE2: RBD spike and (c)-(d) oACE2: RBD spike at 200 ns. ACE2 receptor is marked in pink, RBD spike in cyan and the contact residues are space-filled and highlighted in yellow. (a) Snapshot of hACE2: RBD spike at 200 ns time span.
(b) The region of interaction is shown separately. Green dotted line indicates hydrogen bonding and brown dotted line denotes electrostatic interactions. The residues are labeled in green.
(c) Snapshot of oACE2: RBD spike at 200 ns time span.
(d) Close-up view of the region of interactions. Yellow dotted line denotes hydrophobic interaction. Rest of the color demarcations is same as (b).
Lys31 of oACE2 participates in hydrogen bonding with RBD spike till 30 ns, after which such intermolecular h-bond interaction disappears from the region. However, Lys31of hACE2 forms consistent intermolecular hydrogen bonding interactions till the end of the simulation. Glu35 of both hACE2 and oACE2 forms h-bond with RBD till the end of the simulation. In oACE2 upto 4 h-bonds were found after 120 ns till the end of the simulation. Asp38 of both hACE2 and oACE2 forms h-bond with RBD till the end of the simulation. There appeared two h-bond in oACE2 from 20ns before 140 ns after which some intermittent number of h-bond are found till the end of the simulation. In hACE2, the number of h-bond occurrence is more prominent till the end of the simulation oACE2 at position Thr82 does not take part in the intermolecular h-bond interaction with RBD spike glycoprotein. hACE2 Met82 h-bond is intermittently found between 20 ns to 40 ns. Lys353 in hACE2 does not appear to form h-bond with RBD spike glycoprotein. oACE2 at position Lys353 forms h-bond intermittently, however, such bond is impaired somewhat before the end of simulation (Fig 3).
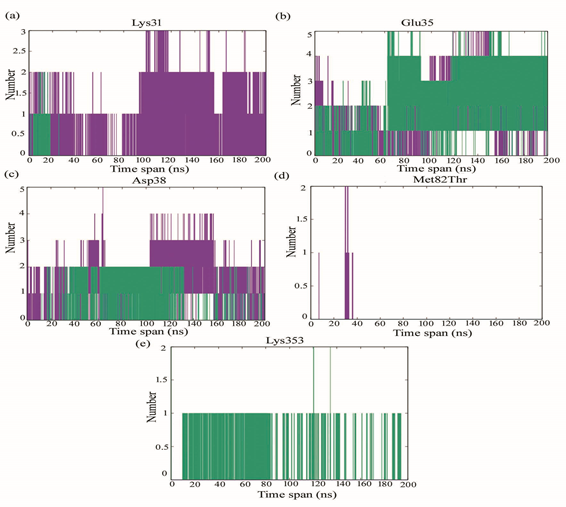
Figure 3: Number of intermolecular hydrogen bonds at (a) Lys31, (b) Glu85, (c) Asp38, (d) Met82Thr and (e) Lys353 of hACE2 (purple) and oACE2 (green) binding with RBD spike glycoprotein of SARS-CoV-2.
The minimum distances between Lys31 in hACE2 and RBD ranges from 0.2-0.4 nm forming stable interaction with RBD spike glycoprotein. Lys31 in oACE2 is found to fluctuate much at various time span of the compressed trajectory. Minimum distance due to Lys31 of oACE2 is found within the range of 0.2 nm at 20ns reflected to form intermolecular h-bond till 20ns. After which the distance is much increased to disrupt any interaction at the region. The minimum distance at position Glu35 for both hACE2 and oACE2 is within the range of 0.2-0.3 nm favorable to form intermolecular h-bond interactions. In Asp38 the minimum distance converged after 100ns for both hACE2 and oACE2. However, after 120ns, the Asp38 in hACE2 is less distantly situated with RBD than oACE2. The minimum distance between Met82 of hACE2 and RBD ranges between 0.2-0.4 nm actively take part in hydrophobic interactions. Thr82 of oACE2 unable to take part in the interaction with RBD being more distantly situated. Lys353 in oACE2 is found to be more closely situated with RBD taking part in interaction whereas; in hACE2 it is much distantly placed to take part in the RBD interactions (Fig. S3).
From the simulated ensemble RMSD, it can be inferred that the oACE2-RBD gets saturated after 100ns time span whereas hACE2-RBD attains saturation after 150ns. The interaction between oACE2 and RBD shows more fluctuations than hACE2-RBD, indicating less stable interaction in oACE2-RBD complex than hACE2-RBD. The binding between hACE2-RBD is strong enough to show less RMSF. The overall compactness (Rg) of the complexes is uniform in nature and binding occurs in both cases. However, more strong interaction is found in hACE2-RBD than oACE2-RBD (Fig. S5).
Interaction energy due to Lennard Jones Potential is more favorable for hACE2::RBD (purple) than oACE2::RBD (green). Interaction energy due to columbic forces is more favorable for hACE2::RBD (purple) than oACE2::RBD (green) (Fig. S4, S5).
4. Discussion
COVID-19 infection is primarily governed by the interaction between human ACE2 and the RBD of the virus SARS-CoV-2. This interaction can also be checked with ACE2 of different animals in order to predict their susceptibility to this virus and the spread thereby. Among the ACE2 of nine farm or pet animals like Felis, Camalus, Bos, Capra, Ovis, Canis, Sus, Oryctolagus and Mus, the binding affinity of Oryctolagus, Felis and Canis ACE2 with the RBD of spike protein was found to be considerably higher (-11.7, -11.9 and -13.8 Kcal/mol, respectively) than ACE2 in remaining livestocks. Burried surface area of Canis ACE2::RBD complex is 2508.8 Å2, which is larger than the ACE2::RBD complex of other livestocks. The ACE2::RBD complexes of Felis and Oryctolagus have buried surface area of 2383.3 Å2 each. So, both binding affinity and buried surface area of complex ACE2::RBD of Felis, Canis and Oryctolagus support the stability in interaction. Overall, the RBD of SARS-CoV-2 is hydrophobic in nature but the ACE2 of Oryctolagus, Felis, Canis are hydrophilic and this assumes to be one of the reasons behind formation of a stable binding complex of ACE2::RBD. Among the three, we have chosen Oryctolagus ACE2::RBD complex for simulation as the ACE2 of this animal belongs to the same clade as human ACE2 and its higher economic importance more in farm industry for fur and meat. Data on simulation performed on Oryctolagus and human ACE2::RBD complex was then compared to get the information of binding efficiency at residue level. In our simulation study with Oryctolagus ACE2::RBD complex, hydrogen bond of Lys31 with RBD does not persist upto the end of the study whereas Lys31 of human ACE2 firmly binds withTyr453 of RBD. Lys31 plays a major role in making a binding platform to RBD. In other hand the stable hydrogen bond formed by the Glu35 of both human and Orytolagus ACE2 with Gln493 and Val445 of RBD respectively, does persist throughout the simulation. Intermittent number of hydrogen bonds is found at the position of Asp38 of Oryctolagus ACE2. Although human Asp38 of ACE2 formed prominent hydrogen bonds till the end of simulation, distance of hydrogen bonds of Oryctolagus also longer than that of the human ACE2. So, compact prominent hydrogen bond is formed at the position of Asp38 of human ACE2. Thr82 is present in Orytolagus ACE2 in place of the Met82 of human ACE2. In human, Met82 forms a hydrogen bond which persists up to 40 ns of simulation. Thr82 of Orytolagus ACE2 does not take part in the formation of any intermolecular hydrogen bond. At the end of the simulation Oryctolagus ACE2 failed to establish interaction with RBD of SARS-CoV-2. In contrast, stable binding affinity is observed between human ACE2 and RBD complex. Anna Z. Mykytyn et.al, 2020[27] has focused on the susceptibility of Orytolagus to SARS-CoV-2, however, detailed molecular interpretation was not provided. It is thus assumed that weak binding affinity of SARS-CoV-2-RBD to ACE2 of Oryctolagus will restrict community transmission than human host.
This study indicates that human to human transmission of SARS-CoV2 is more vulnerable than pet to pet transmission. Although Canis, Felis and Oryctolagus are reportedly infected by SARS-CoV-2, our investigation predicts that overall farm and pets animals will be safe from the vulnerability of SARS-CoV-2 at certain level of infectivity. Humans remain in very close contact with farm animals, pets and laboratory animals for different types of needs. Hence, minimizing the exposures of humans to these animals will further reduce the spillover risks of coronaviruses.
Conflict of interest
The authors declare that they have no conflict of interest.
Declaration
The work has been dedicated to Dr. Suranjita Mitra who was remained inspirational for shaping the idea before her untimely demise in Covid19.
Authors’ Contributions
RB curated, analyzed and interpreted the data. AMG helped in docking and simulation studies. S Mandal and SRB supervised the work. All the authors write, review and edited the manuscript.
References
- Pal M, Berhanu G, Desalegn C, Kandi V. Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2): An Update. Cureus 2 (2020).
- Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nature Medicine 26 (2020): 450-2.
- Han GZ. Pangolins Harbor SARS-CoV-2-Related Coronaviruses. Trends in Microbiology [Internet] 28 (2020): 515-7.
- Shi J, Wen Z, Zhong G, Yang H, Wang C, Huang B, et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS-coronavirus 2. Science 368 (2020): 1016-20.
- Zhou P, Yang X Lou, Wang XG, Hu B, Zhang L, Zhang W, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature [Internet] 579 (2020): 270-3.
- Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367 (2020): 1260-3.
- Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367 (2020): 1444-8.
- Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell [Internet] 181 (2020): 281-292.e6.
- Millet JK, Whittaker GR. Host cell proteases: Critical determinants of coronavirus tropism and pathogenesis. Virus Research [Internet] 202 (2015): 120-34.
- Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor Cell 181 (2020): 271-280.e8.
- Gheblawi M, Wang K, Viveiros A, Nguyen Q, Zhong JC, Turner AJ, et al. Angiotensin-Converting Enzyme 2: SARS-CoV-2 Rece-ptor and Regulator of the Renin-Angiotensin System: Celebrating the 20th Anniversary of the Discovery of ACE2. Circulation Research (2020): 1456-74.
- Guo J, Huang Z, Lin L, Lv J. Coronavirus Disease 2019 (COVID-19) and Cardiovas-cular Disease: A Viewpoint on the Potential Influence of Angiotensin-Converting Enzy-me Inhibitors/Angiotensin Receptor Block-ers on Onset and Severity of Severe Acute Respiratory Syndrome Coronavirus 2 Infect-ion. Journal of the American Heart Associa-tion 9 (2020): e016219.
- Yan R, Zhang Y, Li Y, Xia L, Zhou Q. Structure of dimeric full-length human ACE2 in complex with B 0 AT1 (2020).
- Samavati L, Uhal BD. ACE2, Much More Than Just a Receptor for SARS-COV-2. Frontiers in Cellular and Infection Microbiology 10 (2020): 1-9.
- Vaibhav B. Patel, Jiu-Chang Zhong, Maria B. Grant and GYO. Role of the ACE2/Angiotensin 1-7 axis of the Renin-Angiotensin System in Heart Failure. Circulation Research 118 (2016): 13.
- Wan Y, Shang J, Graham R, Baric RS, Li F. Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. Journal of Virology 94 (2020).
- Park Y, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, et al. The EMBL-EBI search and sequence analysis tools APIs in 2019 F abio 47 (2019): 636-41.
- Robert X, Gouet P. Deciphering key features in protein structures with the new ENDscript server 42 (2014): 320-4.
- Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution 35 (2018): 1547-9.
- Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, et al. SWISS-MODEL: homology modelling of protein structures and complexes (2018): 1-8.
- Willard L, Ranjan A, Zhang H, Monzavi H, Boyko RF, Sykes BD, et al. VADAR: a web server for quantitative evaluation of protein structure quality 31 (2003): 3316-9.
- The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC.
- Zundert GCP Van, Rodrigues JPGLM, Trellet M, Schmitz C, Kastritis PL, Karaca E, et al. SC. Journal of Molecular Biology [Internet] (2015).
- Xue LC, Rodrigues JP, Kastritis PL, Mjj A. Structural bioinformatics PRODIGY: a web server for predicting the binding affinity of protein-protein complexes 2011 (2016): 2014-6.
- Meath WJ, Margoliash DJ, Jhanwar BL, Koide A, Zeiss GD. Interaction Models for Water in Relation to Protein Hydratation. Nature (1981): 331-8.
- Oostenbrink C, Soares TA, Van Der Vegt NFA, Van Gunsteren WF. Validation of the 53A6 GROMOS force field. European Biophysics Journal 34 (2005): 273-84.
- Mykytyn AAZ, Lamers MM, Okba NMA, Breugem TI, Doel PB Van Den, Run P Van, et al. Susceptibility of rabbits to SARS-CoV-2 Affiliations (2020): 1-17.
