Generation and Characterization of A Carboplatin Resistant Ovarian Cancer Model
Article Information
Shalini N Swamy1, Sandeep Kumar S1, Devaraj VR2, Ramesh Gawari1*
1Department of Biochemistry, Kidwai Memorial Institute of Oncology, Bangalore, India
2Department of Biochemistry, Bengaluru Central University, Bangalore, India
*Corresponding Author: Dr. Ramesh Gawari, Department of Biochemistry, Kidwai Memorial Institute of Oncology, Bangalore, India
Received: 01 May 2020; Accepted: 19 May 2020; Published: 01 July 2020
Citation: Shalini N Swamy, Sandeep Kumar S, Devaraj VR, Ramesh Gawari. Generation and Characterization of A Carboplatin Resistant Ovarian Cancer Model. Journal of Cancer Science and Clinical Therapeutics 4 (2020): 144-153.
View / Download Pdf Share at FacebookAbstract
Acquisition of drug resistance is one of the major hurdles in the treatment of several cancers including ovarian carcinoma. There exist very few models that aid in understanding the mechanism of drug resistance in cancer cells. Carboplatin is administered as a first line chemotherapeutic drug in the treatment of ovarian cancer. Majority of ovarian cancer patients respond well to chemotherapy in the initial stages, but 60-70% report recurrence post therapy. The underlying molecular mechanisms contributing to resistance to therapy remain elusive. Establishing a clinically relevant in vitro tumor model that could closely mimic the drug resistance pattern in the patient would provide an opportunity to understand drug resistance. The present study focuses on generating a clinically relevant drug resistant model of ovarian cancer.
Keywords
<p>Drug resistance; Carboplatin; Ovarian cancer; ABCB1; Ku80; Ku70</p>
Drug resistance articles, Carboplatin articles, Ovarian cancer articles, ABCB1 articles, Ku80 articles, Ku70 articles
Drug resistance articles Drug resistance Research articles Drug resistance review articles Drug resistance PubMed articles Drug resistance PubMed Central articles Drug resistance 2023 articles Drug resistance 2024 articles Drug resistance Scopus articles Drug resistance impact factor journals Drug resistance Scopus journals Drug resistance PubMed journals Drug resistance medical journals Drug resistance free journals Drug resistance best journals Drug resistance top journals Drug resistance free medical journals Drug resistance famous journals Drug resistance Google Scholar indexed journals Carboplatin articles Carboplatin Research articles Carboplatin review articles Carboplatin PubMed articles Carboplatin PubMed Central articles Carboplatin 2023 articles Carboplatin 2024 articles Carboplatin Scopus articles Carboplatin impact factor journals Carboplatin Scopus journals Carboplatin PubMed journals Carboplatin medical journals Carboplatin free journals Carboplatin best journals Carboplatin top journals Carboplatin free medical journals Carboplatin famous journals Carboplatin Google Scholar indexed journals Ovarian cancer articles Ovarian cancer Research articles Ovarian cancer review articles Ovarian cancer PubMed articles Ovarian cancer PubMed Central articles Ovarian cancer 2023 articles Ovarian cancer 2024 articles Ovarian cancer Scopus articles Ovarian cancer impact factor journals Ovarian cancer Scopus journals Ovarian cancer PubMed journals Ovarian cancer medical journals Ovarian cancer free journals Ovarian cancer best journals Ovarian cancer top journals Ovarian cancer free medical journals Ovarian cancer famous journals Ovarian cancer Google Scholar indexed journals ABCB1 articles ABCB1 Research articles ABCB1 review articles ABCB1 PubMed articles ABCB1 PubMed Central articles ABCB1 2023 articles ABCB1 2024 articles ABCB1 Scopus articles ABCB1 impact factor journals ABCB1 Scopus journals ABCB1 PubMed journals ABCB1 medical journals ABCB1 free journals ABCB1 best journals ABCB1 top journals ABCB1 free medical journals ABCB1 famous journals ABCB1 Google Scholar indexed journals Ku80 articles Ku80 Research articles Ku80 review articles Ku80 PubMed articles Ku80 PubMed Central articles Ku80 2023 articles Ku80 2024 articles Ku80 Scopus articles Ku80 impact factor journals Ku80 Scopus journals Ku80 PubMed journals Ku80 medical journals Ku80 free journals Ku80 best journals Ku80 top journals Ku80 free medical journals Ku80 famous journals Ku80 Google Scholar indexed journals Ku70 articles Ku70 Research articles Ku70 review articles Ku70 PubMed articles Ku70 PubMed Central articles Ku70 2023 articles Ku70 2024 articles Ku70 Scopus articles Ku70 impact factor journals Ku70 Scopus journals Ku70 PubMed journals Ku70 medical journals Ku70 free journals Ku70 best journals Ku70 top journals Ku70 free medical journals Ku70 famous journals Ku70 Google Scholar indexed journals Intravenous articles Intravenous Research articles Intravenous review articles Intravenous PubMed articles Intravenous PubMed Central articles Intravenous 2023 articles Intravenous 2024 articles Intravenous Scopus articles Intravenous impact factor journals Intravenous Scopus journals Intravenous PubMed journals Intravenous medical journals Intravenous free journals Intravenous best journals Intravenous top journals Intravenous free medical journals Intravenous famous journals Intravenous Google Scholar indexed journals Intraperitoneal articles Intraperitoneal Research articles Intraperitoneal review articles Intraperitoneal PubMed articles Intraperitoneal PubMed Central articles Intraperitoneal 2023 articles Intraperitoneal 2024 articles Intraperitoneal Scopus articles Intraperitoneal impact factor journals Intraperitoneal Scopus journals Intraperitoneal PubMed journals Intraperitoneal medical journals Intraperitoneal free journals Intraperitoneal best journals Intraperitoneal top journals Intraperitoneal free medical journals Intraperitoneal famous journals Intraperitoneal Google Scholar indexed journals gynecological malignancies articles gynecological malignancies Research articles gynecological malignancies review articles gynecological malignancies PubMed articles gynecological malignancies PubMed Central articles gynecological malignancies 2023 articles gynecological malignancies 2024 articles gynecological malignancies Scopus articles gynecological malignancies impact factor journals gynecological malignancies Scopus journals gynecological malignancies PubMed journals gynecological malignancies medical journals gynecological malignancies free journals gynecological malignancies best journals gynecological malignancies top journals gynecological malignancies free medical journals gynecological malignancies famous journals gynecological malignancies Google Scholar indexed journals chemotherapeutic drug articles chemotherapeutic drug Research articles chemotherapeutic drug review articles chemotherapeutic drug PubMed articles chemotherapeutic drug PubMed Central articles chemotherapeutic drug 2023 articles chemotherapeutic drug 2024 articles chemotherapeutic drug Scopus articles chemotherapeutic drug impact factor journals chemotherapeutic drug Scopus journals chemotherapeutic drug PubMed journals chemotherapeutic drug medical journals chemotherapeutic drug free journals chemotherapeutic drug best journals chemotherapeutic drug top journals chemotherapeutic drug free medical journals chemotherapeutic drug famous journals chemotherapeutic drug Google Scholar indexed journals
Article Details
Abbreviations:
OC- Ovarian cancer; IV- Intravenous; IP- Intraperitoneal; CR- Carboplatin Resistant; IC- Inhibitory concentration; DSB- Double strand break; RQ: Relative quantification
1. Introduction
Ovarian cancer (Oc) is one of the most lethal gynecological malignancies afflicting women. It is generally detected in the late stages as it remains asymptomatic in the early stages. The five year survival rate for cancers detected in the late stages is about 26% as against 93% when detected in early stages [1]. The first-line of treatment for advanced OC is cytoreductive surgery to eliminate the bulk of tumors followed by intravenous (IV) or intraperitoneal (IP) administration of carboplatin in combination with taxane. Although most patients initially respond to treatment, the disease recurs in almost 60-70% of patients leading to high mortality [2, 3]. The high mortality rate of ovarian cancer is attributed to both innate and acquired drug resistance [4]. Yet, the underlying molecular basis of therapeutic resistance still remains elusive. Understanding the molecular basis for therapeutic resistance will facilitate the identification of actionable targets and development of new personalized and/or combination therapies for cancer patients. Therefore, there is a need to generate experimental models which are similar to the resistant cells in the patients and thereby aid in understanding the acquisition of carboplatin resistance in ovarian cancer.
Till date, there are no carboplatin resistant ovarian cancer cell lines available or established that could serve as clinically relevant models of OC drug resistance. In this work, we describe the generation of a clinically relevant carboplatin resistant (CR) cell line for ovarian cancer.
2. Methods
2.1 Determination of cell viability and IC50 value of Carboplatin2 x 104 cells were plated in 96 well plates and after 12 h of seeding the cells were treated with various concentrations of carboplatin (0.0 mM, 0.05 mM, 0.10 mM, 0.15 mM, 0.20 mM, 0.25 mM) for 48 hours. After 48 h of drug exposure, MTT (0.5 mg/ml) was added to each well and incubated for 4h until farmazon crystals were formed. The crystals were dissolved using DMSO and the absorbance was read at 570 nm. Cell viability is expressed as percentage of absorbance of drug treated cells relative to that of the untreated. All the assays were performed in triplicates and repeated three times.
2.2 Establishment of drug resistant cells
OVCAR3 cells are one of the widely used in vitro models to study ovarian cancer biology. In order to establish a clinically relevant CR model of OVCAR3 cells, a pulse selection strategy was adopted. OVCAR-3 cells were seeded at density of 1 x 105 cells to form a monolayer. 12 h after seeding, the cells were pulse treated with sub-lethal concentration (30 uM) of carboplatin for 3 hours. After 3hours of treatment, drug containing medium was withdrawn and replenished with drug free complete medium and cultured till the cells reached 75% confluence. Once confluent the cells were seeded in the same density as mentioned previously and the drug regimen was continued in the same manner for 6 cycles (Figure 1). After the drug resistant clone is established, the fold resistance is calculated as a ratio of IC50 of CR cells: Ic50 of parent cells.
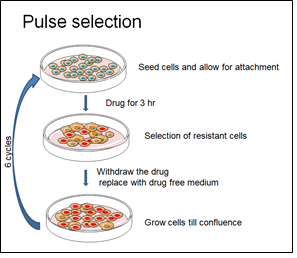
Figure 1: Schematic representation of the methodology adapted to establish drug resistance.
2.3 Gene Expression analysis
RNA was isolated from both parental and drug resistant cells using TRIzol method and reverse transcribed to cDNA using High Capacity cDNA Reverse Transcription Kit (Applied Biosystems ABI, Foster City, CA) following the manufacturers protocol. The cDNA was used as template for gene expression analysis of drug efflux gene ABCB1. The primer sequences and reaction conditions are su mmarized in Table 1.
|
Stage |
Holding |
Cycling |
Melting curve stage |
|||
|
Temperature |
95ºC |
95ºC |
60ºC |
95ºC |
60ºC |
95ºC |
|
Time |
15mins |
15 sec |
1min |
15sec |
1min |
15sec |
|
Primer sequence |
5’-AAATTGGCTTGACAAGTTGTATATGG-3’ 5’- CACCAGCATCATGAGAGGAAGTC-3’ |
|||||
Table 1: RT-PCR reaction conditions and Primer sequence.
2.4 Characterization of drug resistant cells
The Drug resistant cells were characterized for morphology and expression of DNA repair proteins Ku80 and Ku70 using Immunocytochemistry. Briefly, the cells were fixed using 3.7% paraformaldehyde for 20 mins. Post incubation the cells were washed with PBS and permeabilized with 0.1% Tritox-X100 for 20 mins at room temperature. The cells are then washed thrice with PBS and blocked with 1% BSA (30 mins). The cells are then incubated with primary antibody (diluted with 1% BSA in PBST) overnight at 4ºC. After incubation the primary antibody is removed and the cells are washed three times with PBS. The cells are next incubated with secondary antibody for 1 h at room temperature in dark. The cells are counter stained with DAPI to stain for the DNA and visualized.
The acquired images were analyzed using Image J software. The integrated density and grey values of were obtained and the corrected total cell fluorescence (CTCF) was calculated as CTCF = Integrated Density – (Area of selected cell X Mean fluorescence of background reading. Data is expressed as means ± SD and p value is calculated.
2.5 Statistical analysis
Statistical analysis was performed using student’s t-test and a p- value< 0.05 was considered to be statistically significant.
3. Results
3.1 Determination of IC50 of carboplatin in parental cells
OVCAR-3 cells were chosen as the parent cells for the current study as this cell line is derived from the papillary adenocarcinoma origin and over 80% of the patients with ovarian carcinoma fall in this class. IC50 value of carboplatin for parental OVCAR-3 cells was determined by MTT assay using varying concentrations of carboplatin. Effect of each dose of drug was assessed in triplicates and the assay was repeated thrice. The IC50 value of carboplatin was determined to be 84.37µM (Figure 2).
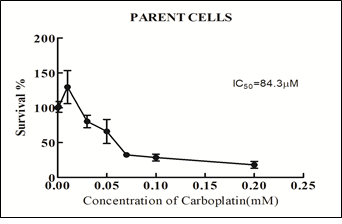
Figure 2: Determination of IC50 in parental cells by MTT assay.
3.2 Generation of Carboplatin resistant cells
The cells were seeded in tissue culture dishes and exposed to sub lethal (30 uM) concentration of carboplatin for 3 hours (pulse selection strategy). Following the three-hour drug exposure, a population of cells detached from the culture dish and began to float. These cells were removed and the cells were replenished with fresh drug free medium and allowed to grow to at least, 75% confluence. It was noteworthy that these cells that remained adherent to the dish after the 3 hour drug exposure doubled slower and appeared larger than the parental counterpart.
The drug treatment was done for a total of 6 cycles. Post 6 cycles of drug exposure, the viability of these cells was assayed to determine the IC50 value. The IC50 value of the carboplatin resistant clones was determined to be 516.2 µM and the fold resistance (IC50 of CR cells/IC50 of parent cell) was found to be 6.34 (Figure 3).
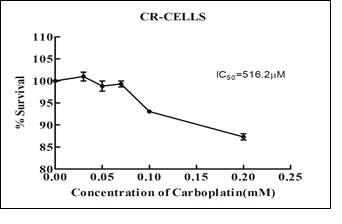
Figure 3: Determination of IC50 of CR- cells after 6 cycles of drug treatment assayed by MTT assay.
3.3 Morphological variations between parental and drug resistant cells
The morphology of both parental and drug resistant cells was monitored during administration of drug. After the treatment of the cells with the initial dosage, we observed that many of the cells during the recovery phase (cells replenished with drug free medium) detached from the tissue culture plastic (8-12th day) and appeared round. However these cells were pelleted and seeded back into a new culture dish and these cells adhered back to the culture dish, indicating its survival strength during acquisition of drug resistance (Figure 4). After the fourth cycle a few cells began to form cellular aggregates in the culture similar to the aggregates that form in the recurrent ovarian cancer patients indicating a close similarity with the clinical setting (Figure 5). The drug resistant cells appeared larger with bigger nucleus that their parent counterpart (Figure 7a and c).
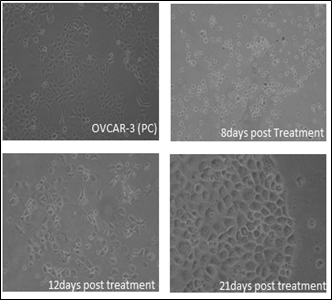
Figure 4: Representative phase contrast images of cell acquired at different time points during drug dosing.
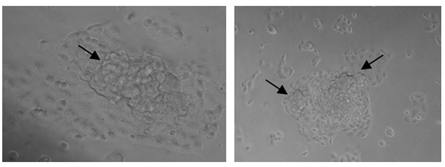
Figure 5: Phase contrast images of CR-cells after 4th cycle of drug dosing depicting the formation of aggregates (black arrow).
3.4 Upregulation of ABCB1 gene
In order to validate the CR-resistant clone that was established, a gene expression analysis of ABCB1 gene, a classical marker that is found to be upregulated in drug resistance was assayed using RT-PCR. The expression of ABCB1 was 3.57 folds upregulated in the resistant cells compared to the parental cells. This confirms the generation of a carboplatin resistant clone (Figure 6).
|
Sample |
RQ value |
|
Parent |
1.0 |
|
CR-cells |
3.571838 |
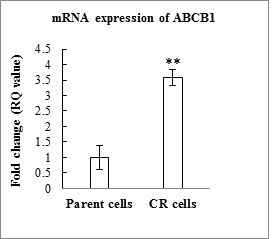
Figure 6: Graphical representation of the mRNA expression of CR-cells and parent cells (p=0.0052).
3.5 Increased expression of DSB repairs proteins
Furthermore, DNA double stranded break repair proteins Ku80 and Ku70 were assayed by I mMunocytochemistry and the mean fluorescent intensity was measured. The mean fluorescent intensity of both Ku80 and Ku70 were higher in the resistant cells as compared to the parental cells. The intensity of Ku80 was found to be statistically significant in CR- cells.
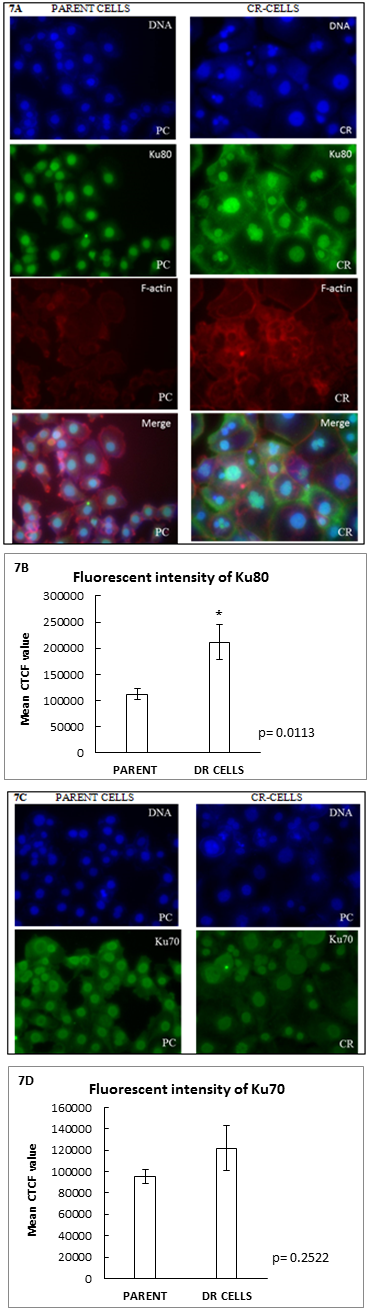
Figure 7: (A and C): Immunofluorescence staining of Ku80 and Ku 70 in parent and CR-cells. The panel of images depict that the CR cells are larger and have bigger nucleus compared to parental cells. (B and D): Graphical representation of mean CTCF value of Ku80 and Ku70 in parental and CR-cells respectively. (Scale: 100µm).
4. Discussion
Carboplatin is considered as a first line drug in the treatment of ovarian cancer. Elevated DNA repair mechanism, increased efflux of drug, and intracellular detoxification mechanisms are known to contribute to the development of chemoresistance in ovarian cancer patients. In the current study, we have generated a carboplatin resistant cell line which mimics the clinical scenario. Clinically relevant drug resistant cells are generated by closely mimicking the conditions that the cancer patients experience during chemotherapy [5]. Such resistant clone/cell lines display a two- to eight-fold resistance compared to their parental cell line. Dosage of drug administered is low, and a pulsed treatment strategy is used where the cells are allowed to recover in drug-free media. The pulse selection strategy produces more stable clones.
The resistant cells generated in this study are larger than their parental counterpart and divide rather slowly similar to the resistant cells that remain within the patient. Such slow cycling phenotypes have been identified in several cancer types and are known to confer a survival advantage within the tumor environment [6-9]. Like in most drug resistant tumors, the mRNA level of the drug transporter protein ABCB1 was also found to be upregulated in CR cells and several studies have reported that ABCB1 up regulation is implicated in poor overall survival and prognosis of ovarian cancer [10-13]. We have further shown that the expression of Ku80, a protein encoded by the XRCC5 gene, involved in DSB repair, is found to be higher in the resistant clone. The higher expression of DSB repair genes are known to contribute to several aspects of tumor progression including drug resistance [14-16].
The generation of carboplatin resistant cells described herein can be used to generate specific drug resistant model that could be a functional tool to explore novel, and relevant molecular and cellular mechanisms of chemo-resistance, which in turn, help design new treatment strategies for highly resistant cancers.
Acknowledgements
We thank Dr. Ramray Bhat and his lab member (Indian Institute of Science, Bangalore) for extending scientific support. We thank Dr.Vishal Singh, Department of Pharmacology, Physiology and Neuroscience, Rutgers New Jersey Medical School, USA, for his kind help in statistical analysis. The authors acknowledge Bristol-Myers Squibb for sponsoring the cell culture facility at the Department of Biochemistry, Central College, Bangalore.
Conflict of Interest
The authors declare no conflict of interest.
Funding
The authors acknowledge Rajiv Gandhi University of Health Sciences, Bangalore for the Research grant (Grant Number: RI004, 2016). SKS is the recipient of Senior Research Fellowship, Council of Scientific and Industrial Research (09/999/0002/2016-EMR-I).
References
- Sandeep Kumar S, Shalini N Swamy, C S Premalatha et al. Aberrant Promoter Hypermethylation of RASSF1a and BRCA1 in cell free circulating tumor DNA serves as a biomarker of ovarian carcinoma. Asian Pacific Journal of Cancer prevention 20 (2019): 3001-3005.
- Ozols RF. Treatment goals in ovarian cancer. International Journal of Gynecologic Cancer 15 (2005): 3-11.
- Rubin SC, Randall TC, Armstrong KA, et al. Ten-year follow-up of ovarian cancer patients after second-look laparotomy with negative findings. Obstetrics & Gynecology 93 (1999): 21-24.
- Sousa GF, Wlodarczyk SR, Monteiro G. Carboplatin: molecular mechanisms of action associated with chemoresistance. Brazilian Journal of Pharmaceutical Sciences 50 (2014): 693-701.
- McDermott M, Eustace A, Busschots S, et al. In vitro development of chemotherapy and targeted therapy drug-resistant cancer cell lines: a practical guide with case studies. Frontiers in oncology 6 (2014): 40.
- Ahn A, Chatterjee A, Eccles MR. The slow cycling phenotype: a growing problem for treatment resistance in melanoma. Molecular cancer therapeutics 16 (2017): 1002-1009.
- Moore N, Houghton J, Lyle S. Slow-cycling therapy-resistant cancer cells. Stem cells and development 21 (2012):1822-1830.
- Kondoh E, Mori S, Yamaguchi K, et al. Targeting slow-proliferating ovarian cancer cells. International journal of cancer 126 (2010): 2448-2456.
- Hamis S, Nithiarasu P, Powathil GG. What does not kill a tumour may make it stronger: In silico insights into chemotherapeutic drug resistance. Journal of theoretical biology 454 (2018): 253-267.
- Sun S, Cai J, Yang Q, et al. Prognostic value and implication for chemotherapy treatment of ABCB1 in epithelial ovarian cancer: a meta-analysis. PloS one 11 (2016).
- Vaidyanathan A, Sawers L, Gannon AL, et al. ABCB1 (MDR1) induction defines a common resistance mechanism in paclitaxel-and olaparib-resistant ovarian cancer cells. British journal of cancer 115 (2016): 431-441.
- Genovese I, Ilari A, Assaraf YG, et al. Not only P-glycoprotein: amplification of the ABCB1-containing chromosome region 7q21 confers multidrug resistance upon cancer cells by coordinated overexpression of an assortment of resistance-related proteins. Drug Resistance Updates 32 (2017): 23-46.
- Wang X, Zhang H, Chen X. Drug resistance and combating drug resistance in cancer. Cancer Drug Resist 2 (2019): 141-160.
- Liu ZH, Wang N, Wang FQ, et al. High expression of XRCC5 is associated with metastasis through Wnt signaling pathway and predicts poor prognosis in patients with hepatocellular carcinoma. European review for medical and pharmacological sciences 23(2019): 7835-7847.
- Kalra RS, Bapat SA. Enhanced levels of double-strand DNA break repair proteins protect ovarian cancer cells against genotoxic stress-induced apoptosis. Journal of ovarian research 6 (2013): 66.
- Wang S, Wang Z, Yang YU, et al. Overexpression of Ku80 correlates with aggressive clinicopathological features and adverse prognosis in esophageal squamous cell carcinoma. Oncology letters 10 (2015): 2705-2712.
