A Cell-free Infection System to Study Translation, Replication and Phage-particle Production during Infection of E. coli By Bacteriophage Qβ
Article Information
Pernille Skov Rasmussen, Charlotte Rohde Knudsen*
1Department of Molecular Biology & Genetics, University of Aarhus, Universitetsbyen 81, DK-8000 Aarhus C, Denmark
*Corresponding author: Charlotte Rohde Knudsen, Department of Molecular Biology & Genetics, University of Aarhus, Universitetsbyen 81, DK-8000 Aarhus C, Denmark.
Received: 24 March 2022; Accepted: 30 March 2022; Published: 26 May 2022
Citation:
Pernille Skov Rasmussen, Charlotte Rohde Knudsen. A Cell-free Infection System to Study Translation, Replication and Phage-particle Production during Infection of E. coli by Bacteriophage Qβ. Journal of Biotechnology and Biomedicine 5 (2022): 94-116.
View / Download Pdf Share at FacebookAbstract
Background: Viruses infect all kingdoms of life, and new species are continuously being discovered. The single-stranded (+)-RNA viruses comprise the largest group of viruses, which includes pathogens such as Dengue virus, Corona virus and West Nile virus. Also, the simple bacteriophage Qβ belongs to this group of viruses. Studies of the mechanism of Qβ infection can increase our general understanding of the single-stranded (+)-RNA viruses, which can be exploited in the pursuit to treat and prevent diseases caused by pathogenic (+)-RNA viruses.
Methods: In this study, we have analysed the production of infectious Qβ phage particles in three different cell-free infection systems upon addition of the Qβ genome as a template. The cell-free infection systems were based on cell-free protein expression systems: two commercial systems and one custom-made system. We studied the course of infection by analysing the production of viral RNA, proteins and phage particles produced in the cell-free reactions. The replication of the viral RNA was determined by RT-PCR, while the translation of the viral proteins was examined by radiolabelling, and the production of infectious phage particles was evaluated by double-layered plaque assays.
Results: Bacteriophage Qβ was found to replicate in two of the three tested cell-free infection systems. Specifically, the viral RNA was replicated, the viral proteins were translated, and infectious phage particles were produced in the cell-free infection systems. The pattern of translation regulation of the viral proteins appeared similar to in vivo infection. Infectious Qβ phage particles were produced at yields of 2.5 × 105 PFU/μL reaction and 2.5 × 103 PFU/μL reaction in the commercial and custom-made system, respectively. Importantly, intact Qβ phage particles were shown not to replicate in the cell-free infection syst
Keywords
<p>Single-stranded (+)-RNA virus; Bacteriophage Qβ; Host factor Q; Cell-free synthetic biology; Cell-free protein expression; Cell-free infection system; Self-assembly; RT-PCR; [14C] labelling; Double-layered plaque assay</p>
Article Details
Abbreviations:
CFI: Cell-free infection; CFPS: Cell-free protein synthesis; Hfq: Host factor Q; PFU: Plaque-forming units; RdRP: RNA-dependent RNA polymerase; RT-PCR: Reverse transcription polymerase chain reaction
1. Background
Viruses are intracellular parasites that infect all three domains of life. They rely on their target cell for molecular building blocks, energy sources and host factors to replicate, since viruses are gene poor compared to their host [1]. Virus particles have a nucleic acid genome and may also carry a few viral proteins, which are enclosed in a capsid of viral proteins or an envelope comprised of a small part of the host cell membrane. The genome consists of either RNA or DNA and can be single- or double-stranded, linear or circular. Viruses have been divided into seven groups in the Baltimore classification, which is based on the nature of their genome and the replication hereof [2]. The largest group is the single-stranded, positive sense (+)-RNA viruses, which comprises more than a third of all viruses. Common for these viruses is that their genomes resemble mRNA and act directly as a template for translation upon infection. The group of (+)-RNA viruses includes a number of pathogenic viruses such as Polio virus, Hepatitis A and C, Dengue virus and Severe Acute Respiratory Syndrome coronavirus [3]. The (+)-RNA viruses have been used in the development of vaccines, as imaging tools and in virus inactivation studies [4, 5]. This group of (+)-RNA viruses also includes numerous bacteriophages of which most are poorly characterised. The number of sequenced genomes of single-stranded bacteriophages has gone from ~30 to more than 15,000 in the past five years, all with potential therapeutic and biotechnological relevance [7,8]. The simple bacteriophage Qβ is an example of a very well-characterised single-stranded (+)-RNA virus, that has been frequently used as a model to study more complex pathogenic (+)-RNA viruses [7]. Bacteriophage Qβ infects the gram-negative bacterium Escherichia coli (E. coli), but it only targets so-called male bacteria with F-pili expressed on the surface. The Qβ genome is only 4217 nucleotides long and encodes a maturation protein, a coat protein, a read-through protein and a RNA-dependent RNA polymerase (RdRP; known as the β-subunit) [7,8]. The read-through protein is occasionally produced as a result of leaky termination during translation of the coat protein [9]. The icosahedral Qβ capsid enclosing the (+)-RNA genome is composed of 176 copies of the coat protein, 3-5 read-through proteins and one molecule of the maturation protein [10, 11]. During the first step of the Qβ life cycle, the phage particle adsorbs to the surface of E. coli via contact between the maturation protein on the viral surface and the bacterial F-pilus [12, 13]. This interaction promotes the release of the Qβ (+)-RNA genome into the host cell via a poorly understood mechanism, which may rely on the retraction of the F-pilus to pull the complex of the maturation protein and the Qβ (+)-RNA into the host cell [13, 14]. Next, the Qβ (+)-RNA is translated by host ribosomes in a tightly controlled process [15]. Initially, the coat protein is translated, which causes a rearrangement of the secondary structure of the (+)-RNA to enable the subsequent translation of the replicase gene and the maturation gene [16]. The coat proteins form dimers that bind to the operator region of the replicase gene, thereby blocking the binding of ribosomes and consequently abolishing translation [17]. After 45 minutes of infection, approximately 5 × 105 molecules of coat proteins are accumulated, whereas the other proteins are produced in much lower amounts e.g. only approximately 450 molecules of the β-subunit are produced [10]. The Qβ (+)-RNA is replicated via formation of a complementary (–)-strand intermediate as soon as the β-subunit is produced. The active Qβ replicase complex is composed of the viral β-subunit and the host proteins EF-Tu, EF-Ts and ribosomal protein S1 [18]. Also host factor Q (Hfq) assists the Qβ replicase complex by recognising the 3’ end of the Qβ (+)-RNA [19], and ensuring a higher affinity for Qβ (+)-RNA than for host mRNAs [20]. In the uninfected cell, Hfq acts as a global regulatory protein that controls the expression of many mRNAs by facilitating interactions between small regulatory RNAs and mRNA molecules [21]. Late in the infection cycle, the (+)-RNA virus particles are assembled. The maturation proteins accumulate in the cell wall of the host cell, which results in cell lysis and release of the progeny viruses [22].
The study of the kinetics of viral infections in vivo relies on the ability to synchronise the infection [23, 24], which depends on the infectivity and cellular conditions of the individual cells. Common synchronisation methods include inoculation at 4°C and starvation of which the first approach lowers the infectivity of the cells, and both approaches may affect other cellular processes [24, 25]. Magnetofection is an alternative method of synchronisation of in vivo infections, which however, is technically demanding [26]. Cell-Free Protein Synthesis (CFPS) systems allow the setup of complex systems, where reaction networks including viral infections can be studied. Upon addition of a viral genome to a host cell extract containing the relevant host components, a Cell-Free Infection (CFI) system is established that results in the production of viral particles. CFI systems provide an attractive alternative of synchronising a viral infection of interest. Importantly, CFI systems allow the study of the role of essential host proteins in viral replication, without considering the survival of host cells [27]. Furthermore, viral infections can be studied under different environmental conditions e.g. the cell-extract can be prepared from cells grown under stressed or other potentially interesting conditions, since a cell-extract made from stressed cells can still support protein synthesis [28]. In this study, we test the ability of three different cell-free protein synthesis systems to support the production of infectious phage particles by directly adding the Qβ RNA genome or a cDNA copy hereof to the cell extract, thereby generating a CFI system. We show that the Qβ phage can replicate in two out of three tested CFI systems. Specifically, the Qβ (+)-RNA is translated and replicated, and infectious progeny phage particles are successfully assembled. Furthermore, we show that intact Qβ phage particles cannot replicate in a CFI system.
2. Methods
2.1. Preparation of viral templates
The pQβ7 plasmid is an infectious plasmid encoding a full-length cDNA copy of the Qβ genome [29]. The pQβ7 plasmid holds an ampicillin resistance gene. The plasmid was purified using a Midiprep kit (Nucleobond Xtra Midiprep, Macherey-Nagel) followed by phenol/chloroform extraction and ethanol precipitation. The purified pQβ7 plasmids were used as a template in two different ways: directly as a template in CFI systems or as a template for in vitro RNA transcription of Qβ (+)-RNA, which subsequently served as the template in CFI systems. In the latter case, the pQβ7 plasmid was linearized using SmaI (Thermo Scientific) to ensure well-defined 3’-ends of the resulting transcripts, while the 5’-ends are dictated by the upstream T7 promoter. The linearized DNA was purified by phenol/chloroform extraction and ethanol precipitation. Qβ (+)-RNA was produced by the “TranscriptAid T7 High Yield Transcription Kit” (Thermo Scientific) using the linearized pQβ7 as a template. The RNA products were purified by phenol/chloroform extraction followed by ethanol precipitation and the yield and homogeneity of the RNA was determined by 1% agarose gel electrophoresis.
2.2. Isolation of Qβ phage particles
- coli HB101 cells were transformed with pQβ7 and grown in 100 mL LB medium for 48 hours at 37°C with shaking. The culture was centrifuged at 12,000 ×g for 30 minutes at 4°C, and the resulting supernatant containing the Qβ phage particles was filtered through a 0.45 μm sterile filter followed by filtration through a 0.2 μm sterile filter. The Qβ phage particles were precipitated for 48 hours in a solution with a final concentration of 9% PEG 6000 and 0.45 M NaCl, and subsequently centrifuged at 15,000 ×g for 30 minutes at 4°C. The pellet was dissolved in 3 mL 40 mM Tris/HCl pH 7.6, 20 mM MgCl2 and 200 mM NaCl followed by dialysis overnight against the same buffer. The phage particle solution was stored in 33% glycerol at -20°C.
2.3. Purification of Hfq
BL21(DE3) was transformed with pTYB11-6 (New England Biolabs) encoding Hfq with a N-terminal tag consisting of a chitin-binding domain embedded in an intein tag for ease of purification [30]. A 2-L culture was inoculated with 1.25% overnight culture of the transformed cells and grown in auto-induction medium (10 g/L peptone, 5 g/L yeast extract, 10 g/L NaCl, 25 mM Na2HPO4, 25 mM KH2PO4, 10 mM NH4Cl, 2 mM MgSO4, 0.5% glycerol, 0.05% D-(+)-glucose, 0.2% L-(+)-lactose, 100 μg/mL ampicillin) at 37°C overnight. The cells were harvested and resuspended in column buffer (20 mM Tris/HCl pH 8, 500 mM NaCl, 1 mM EDTA and 1 mM PMSF pr. gram cells) and subsequently lysed with lysozyme (0.5 mg/g cells) and deoxycholate (0.1 mL/g cells). The lysed cells were DNase treated (0.5 mg/mL, Roche) and centrifuged (4,500 rpm, 40 minutes). The supernatant was applied to a chitin column (New England Biolabs) equilibrated with column buffer followed by washing overnight with the same buffer at 4°C. On the following day, the column was flushed with flush buffer (20 mM Tris/HCl pH 8, 500 mM NaCl, 1 mM EDTA and 50 mM DTT), the flow was stopped, and the column was incubated for 24 hours at 4°C and for 24 hours at room temperature. Hfq liberated by intein splicing was eluted with column buffer and dialysed overnight at 4°C against dialysis buffer (20 mM Tris-HCl, 200 mM NaCl). The protein concentration was determined by the Bradford procedure (Bio-Rad).
2.4. Cell-free infection systems
The ability of the Qβ virus to replicate in a bacterial cell-free system was tested in three different cell-free protein expression systems, which were either of commercial origin or homemade. The Qβ genome was used as a template either as a plasmid-borne cDNA copy or as an in vitro transcribed (+)-RNA, which were directly added to the systems. The “PURExpress® In Vitro Protein Synthesis” (PURE) kit (New England Biolabs) contains all the purified E. coli components required for cell-free translation. Additionally, it contains T7 RNA polymerase to enable transcription from a DNA template driven by a T7 promoter. 25-μL reactions were set up following the instructions from the manufacturer with intact pQβ7 plasmid (0.8 μg) or Qβ (+)-RNA (6 μg) as templates. Since Hfq was not expected to be part of the system, this host protein was either added (4.4 μg) in 10-1,000 times molar excess over added genomes or left out to study its importance. The reactions were incubated at 37°C for 4 hours at 160 rpm. The “S30 T7 High-Yield Protein Expression System” (S30 T7) (Promega) is a kit containing an E. coli extract, which has been subject to a S30 centrifugation step. The extract contains T7 RNA polymerase. CFI reactions of 25 μL were set up using this kit with pQβ7 plasmid (0.8-3.2 μg), Qβ (+)-RNA (3-12 μg) or control DNA (provided by the manufacturer) as templates. The protocol from the manufacturer was followed and the reactions were incubated at 25°C for 8-16 hours or at 37°C for 3 hours at 160 rpm. The ability of intact Qβ phage particles to replicate in a CFI system was tested using the S30 T7 kit. Reactions containing ~50 or ~200 Qβ phage particles instead of the DNA or RNA template were prepared. A separate reaction with 1.6 μg pQβ7 was included as a positive control. The reactions were incubated at 25°C for 16 hours at 160 rpm. A CFPS system was made by growing and processing BL12(DE3) E. coli into a functional S30 T7 extract following the protocol established by Krinsky et al. [31]. The extract contains T7 RNA polymerase, since the strain contains the λDE3 lysogen that carries the gene for T7 RNA polymerase. The buffers and solutions were prepared as described by Krinsky et al. [31]. The aromatic amino acids were dissolved by adding a small amount of NaOH, and undissolved amino acids were removed by filtration (0.45 μm). E. coli BL21(DE3) cells were grown as described by Krinsky et al. [31]. The cells were harvested, and the pellet was resuspended in 30 mL lysate buffer (10 mM Tris-acetate pH 7.4, 14 mM magnesium acetate, 60 mM potassium acetate, 1 mM DTT, 7.2 mM 2-mercaptoethanol). The cells were broken by passaging through a High-Pressure Homogeniser at 15,000 psi twice. 50-μL reactions added 40 U “Ribolock RNase inhibitor” (Thermo Scientific) were set up with Qβ (+)-RNA (24 μg) or pQβ7 (0.6 μg) as templates. Control reactions with a pET vector encoding a 10 kDa protein (0.5 μg) or without a template were included. The reactions were incubated either at 25°C for 5-16 hours or 37°C for 16 hours at 160 rpm.
2.5. Analysis of phage production
The production of infectious Qβ phages in the CFI systems was evaluated with a double-layered plaque assay [32]. Purified Qβ phages were included as a positive control, and top agar, uninoculated overnight culture, LB medium and H2O were included as negative controls to assess potential sources of contamination. The indicator E. coli strain K603 (Coli Genetic Stock Center, Yale University; F+, thr-1, araC14, leuB6(Am), lacY1, glnX44(AS), galK2(Oc), galT22, λ-, ΔtrpE63, xylA5, mtl-1, thiE1, F1-10(Tn10)) was grown on a LB agar plate with tetracycline overnight at 37°C. 250 μL of the overnight culture was added to 25 mL LB media (37°C) with tetracycline and incubated at 37°C, 160 rpm until OD600≈0.6. The cells were harvested by centrifugation (5,000 rpm, 5°C, 15 minutes), and the pellet was resuspended in 5 mL LB media added tetracycline. LB agar plates were made with added supplements (2 mM CaCl2, 0.1% glucose, 10 μg/mL thiamine). Autoclaved top agar (10 g/L peptone, 5 g/L yeast extract, 10 g/L NaCl, 5 g/L agar, 5% glycerol) was mixed with supplements and tetracycline and aliquoted into sterile 4-mL tubes placed at 60°C on a heating block. The top agar was cooled briefly before addition of 100 μL of the freshly grown K603 E. coli cells followed by pouring on the LB agar plates. The CFI reactions were analysed undiluted and in dilutions in the range from 10-2 to 10-10, while only the undiluted control reaction without template was analysed, as were all the other negative controls. A positive control with purified Qβ phages was included. 10 μL of each dilution was transferred to 190 μL LB media and spread on the solidified top agar. The plates were incubated at 37°C overnight. On the following day, plaque-forming units (PFU) were counted on the plates, where density allowed a sound determination of the number of PFU, and the dilution was taken into account resulting in the total PFU (PFUtotal) in the sample. The average number of the PFUtotal was calculated, when countable numbers of plaques appeared on several plates representing different dilutions of the same reaction.
The concentration of PFU/μL was calculated:
2.6. Analysis of viral protein production
Where indicated, radioactive [14C]-serine (SA 2.75 mCi/mmol) was included in the CFPS reactions following the protocol of Krinsky et al. [31] to allow detection of the produced proteins. The reactions were incubated at 25°C for 16 hours at 160 rpm. The CFPS samples (10 μL) were run on a 4%-20% gradient Mini-PROTEAN TGX Precast Protein Gel (Bio-Rad), which was subsequently fixed in fixer solution (50% ethanol, 10% acetic acid, 3% glycerol) for 30 minutes and dried at 80°C in a gel dryer overnight. The dried gel was placed in a “Molecular Dynamics Storage Phosphor Screen” and exposed for 4 days. The cassette was scanned in a “Typhoon Trio” imager (GE Healthcare). An area of the gel containing a high concentration of free [14C]-serine caught in the front was omitted to achieve higher sensitivity, and the gel was exposed for an additional 6 days and scanned again. The brightness and contrast were optimised equally on the entire blot.
2.7. Analysis of viral RNA replication by RT-PCR
The viral RNA replication in the CFPS system was analysed by reverse transcription polymerase chain reaction (RT-PCR). Total RNA was purified from CFPS reactions added either pQβ7, Qβ (+)-RNA or pET vector as a template and from purified, intact Qβ phages using the “Nucleospin RNA” kit (Macherey Nagel). Additional DNase treatment of the RNA was performed using the “RNase-free DNase I” kit (Thermo Scientific). The purified RNA was analysed on a 1% agarose gel, and the RNA concentration was determined on a “Nanodrop” instrument measuring the absorbance at 260 nm. First-strand synthesis was carried out using the “RevertAid First Strand cDNA Synthesis” kit (Thermo Scientific) using the Qβ_genome_3’UTR primer (Sigma) (Table 1). “Herculase II Fusion DNA Polymerase” (Agilent Technologies) was used in the subsequent PCR reactions. RNA extracted from intact Qβ phages was included as a positive control for the presence of Qβ (+)-RNA, while negative controls without added template, without primer or without reverse transcriptase were included as well. The primers used in the subsequent PCR were Beta-upstream (Sigma) and Qbeta_rev (Sigma) (Table 1). The forward primer binds upstream of the β-subunit gene, while the reverse primer binds within the β-subunit gene (Figure 1). The resulting PCR product has a size of 1036 bp. PCR cycling began with template denaturation at 95°C for 1 min, then 30 cycles of 95°C for 20 sec, 57°C for 20 sec and 72°C for 1 minute and one cycle at 72°C for 5 minutes. The quantity and size of the resulting PCR products were analysed on a 1.5% agarose gel, where 4-μL samples were loaded. The brightness and contrast were optimised equally on the entire gel.
|
Primer |
Tm (°C) |
5’ 3’ |
Application |
|
Beta-upstream |
66 |
CGCGATCCATCCAGTGGTGG |
PCR |
|
Qbeta_rev |
62 |
CCTTAGGTGATCTGAGGTCC |
PCR |
|
Qβ_genome_3’UTR |
59 |
GGGCAAAGCAGATCCCCCTC |
cDNA synthesis |
Table 1: Primers used in RT-PCR. The melting temperature (Tm) is indicated for the primers used for RT-PCR. The primer sequences are shown in the 5’-3’ direction, and their specific applications are provided. All primers were from Sigma.
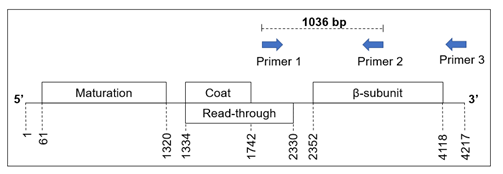
Figure 1: Map of the Qβ genome with binding sites of primers for RT-PCR. The boxes symbolise open reading frames encoding the indicated phage proteins (maturation, coat, read-through and β-subunit). The respective positions of start and stop codons in the Qβ genome are given under the broken lines. The arrows indicate the annealing position and direction of the relevant primers (Table 1). PCR primer 1: Beta-upstream, PCR primer 2: Qbeta_rev and first-strand synthesis primer 3: Qβ_genome_3’UTR. The resulting PCR product has a size of 1036 bp as indicated. The figure is based on information from Klovins et al. [33].
3. Results
In this study, three different CFPS systems were tested for their ability to sustain the production of infectious phage particles and thereby work as CFI systems by adding the viral genome as a template. Two of the tested systems are commercial (the “PURExpress® In Vitro Protein Synthesis” system from New England Biolabs (PURE) and the “S30 T7 High-Yield Protein Expression System” Promega (S30 T7)), while the third system is based on a homemade CFPS system [31]. Table 2 provides an overview of the three systems.
|
System |
Content |
Reactions, |
Reactions, Qβ (+)-RNA |
Hfq |
|
PURE |
Kit with purified proteins from E. coli |
T7 transcription Translation Replication |
Translation Replication
|
Separately added |
|
S30 T7 “S30 T7 High-Yield Protein Expression System” (Promega) |
Kit with E. coli extract |
T7 transcription Translation Replication |
Translation Replication
|
Expected to be present |
|
CFPS Cell-free protein synthesis system (Krinsky et al., 2016)(31) |
E. coli extract from cells grown in the lab |
T7 transcription Translation Replication |
Translation Replication
|
Expected to be present |
Table 2: Overview of the applied cell-free infection (CFI) systems. “Content” indicates the components of the different systems. All systems contain T7 RNA polymerase and can work with either a DNA template (pQβ7) (29), or a RNA template (Qβ (+)-RNA). “Reactions, pQβ7” indicates the processes taking place in the reaction, when pQβ7 is added in the CFI system. Translation and replication are normal steps in the viral life cycle (italics), while T7 transcription of DNA is an additional step that is independent of the viral infection cycle (non-italics) and happens initially, when pQβ7 is used as a template in the CFI systems. “Reactions, Qβ (+)-RNA” indicates the processes involving the Qβ genome during infection, when Qβ (+)-RNA is added in the CFI system. “Hfq” indicates if Hfq is added externally or is expected to be intrinsically present in the CFI system.
3.1. Phage production in commercial protein-expression systems
The two commercial protein-expression systems, PURE and S30 T7, were tested for their ability to support viral infection and cause phage particle production upon addition of the Qβ genome either as an infectious plasmid (pQβ7) [29] or as Qβ (+)-RNA. The PURE system contains purified components necessary for T7 transcription and translation. Two separate reactions were set up with or without Hfq, which is not considered an essential component of protein expression, but is required for Qβ infection [19]. The S30 T7 system is based on an E. coli cell extract that has been prepared by S30 centrifugation. The production of infectious Qβ phages in the commercial CFI systems (PURE and S30 T7) was analysed with a double-layered plaque assay [32]. Purified Qβ phages were included as a positive control, along with the relevant negative controls to monitor contamination. The results show that no infectious Qβ phage particles were produced in either of the PURE systems irrespective of the addition of Hfq, whereas infectious Qβ phage particles were produced by the S30 T7 system with a yield of 3.7 × 104 PFU/μL reaction (Table 3).
|
PFU count 10-2 dilution |
PFU count 10-4 dilution |
PFU count 10-6 dilution |
Yield PFU/μL |
|
|
Positive control Qβ phages |
nd |
nd |
8 |
8 × 105
|
|
PURE +Hfq |
0 |
0 |
0 |
0 |
|
PURE -Hfq |
0 |
0 |
0 |
0 |
|
S30 T7 pQβ7 |
nd |
37 |
0 |
3.7 × 104
|
Table 3: Infectious Qβ virus particles are produced in the S30 T7 system. The double-layered plaque assay was used to analyse the production of phage particles in the PURE and S30 T7 systems. Purified Qβ phages were included in the plaque assay as a positive control (upper row). The PURE system was applied with (+) or without (-) added Hfq. The PURE system was set up with pQβ7 (0.8 μg) and incubated at 37°C for 4 h. The S30 T7 system was set up with pQβ7 DNA (3.2 μg) and incubated at 25°C for 8 h. “PFU count 10-2 dilution”, “PFU count 10-4 dilution” and “PFU count 10-6 dilution” indicate the observed number of plaque-forming units (PFU) on the plates containing the 10-2, 10-4 and 10-6 dilutions, respectively. The number of plaques was counted, where density allowed a sound determination of the number of PFU. “Yield PFU/μL” indicates the number of PFU related to the volume of the CFI reaction. “nd”: not determined due to a high plaque density making the precise counting impossible.
3.2. Intact Qβ phage particles do not replicate in a CFI system
The ability of intact Qβ phage particles to replicate in a CFI system was tested using the S30 T7 system to verify that the observed number of PFU in the CFI systems are products directly derived from the added Qβ genomes. Purified Qβ phage particles were added to the S30 T7 system at two different concentrations (1 or 4 Qβ phages/μL), and the replication of Qβ phages was compared to positive controls with pure Qβ phages analysed with a double-layered plaque assay (Figure 2, Table 4 and Table S1 in Additional file 1). A negative control of the S30 T7 system without added template was included. The positive controls with pure Qβ phages analysed directly a double-layered plaque assay (Table S1 in Additional file 1) resulted in PFU counts that were not considerably lower than the expected number of PFU.
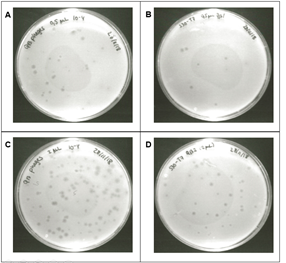
Figure 2: Intact Qβ virus particles do not replicate in the S30 T7 system. A double-layered plaque assay was used to compare the number of PFU obtained after direct plating of pure Qβ phages (Table S1 in Additional file 1) and Qβ phages incubated in the S30 T7 reactions (Table 4). Cleared zones called plaques appear on the surface of a bacterial lawn, where a Qβ phage particle has infected and lysed the surrounding bacteria. The number of plaques was counted, where density allowed a sound determination of the number of PFU. A: ~50 expected purified Qβ phages plated out. B: 10 expected Qβ phages from the S30 T7 system incubated at 25°C overnight. C: ~200 expected purified Qβ phages plated out. D: ~40 expected Qβ phages from the S30 T7 system incubated at 25°C overnight.
Additional file 1.pdf, Table S1. Positive controls included in the double-layered plaque assay. Table S1 in Additional file 1 shows the results of the positive controls with pure Qβ phages analysed with a double-layered plaque assay.
|
PFU count Undiluted |
PFU count 10-2 dilution |
PFU count 10-3 dilution |
PFU count 10-4 dilution |
Yield PFU/μL |
|
|
S30 T7 1 PFU/μL |
8 |
0 |
na |
0 |
0.8 |
|
S30 T7 4 PFU/μL |
38 |
1* |
na |
0 |
3.8 |
|
S30 T7 pQβ7 |
na |
nd |
nd |
~254 |
2.5 × 105 |
|
S30 T7 No template |
0 |
na |
na |
na |
0 |
Table 4: Intact Qβ virus particles do not replicate in the S30 T7 system. Phage particles (1 or 4 PFU/μL) or pQβ7 (1.6 μg) were added to the S30 T7 system to investigate their ability to sustain the production of infectious phage particles upon incubation in the system. A negative control without template was made in parallel. The PFU yield of the reactions was analysed with a double-layered plaque assay. “PFU count undiluted”, “PFU count 10-2 dilution”, “PFU count 10-3 dilution” and “PFU count 10-4 dilution” indicate the observed number of plaque-forming units (PFU) on the plates containing the undiluted sample or the 10-2, 10-3 and 10-4 dilutions, respectively. The number of plaques was counted, where density allowed a sound determination of the number of PFU. *1 PFU is not considered statistically sound. “Yield PFU/μL” indicates the number of PFU related to the volume of the CFI system. “nd”: Not determined due to a high plaque density making the precise counting impossible. “na”: Not analysed.
The expected number of phage particles in the S30 T7 reactions with added Qβ phages were 1 and 4 PFU/μL, respectively, based on the assumption that intact phage particles cannot serve as templates in a CFI system. These numbers are comparable to the observed PFU counts of 0.8 and 3.8 PFU/μL, respectively (Table 4). This confirms that intact Qβ phage particles in the CFI system do not support the production of Qβ phages and justifies that the observed PFU counts reported in Table 3 entirely results from the replication of the added DNA template. The included positive control (Table S1) confirms the functionality of the system with a yield equivalent to the replication of Qβ reported in Table 3.
3.3. Production of infectious Qβ phage particles in the CFPS system
The homemade CFPS system is based on the protocol described by Krinsky et al. [31]. The reaction involves an E. coli cell extract, which has been prepared by S30 centrifugation after cell lysis and removal of debris. Furthermore, amino acids, nucleotides, an energy source and salts are required. The Qβ genome was added to the CFPS system either as an infectious plasmid (pQβ7) or directly as Qβ (+)-RNA. The production of infectious Qβ phage particles by the CFPS system was evaluated with a double-layered plaque assay (Table 5). The results show that infectious Qβ phage particles were successfully produced in the CFPS systems. The CFPS system gave rise to comparable numbers of Qβ phage particles when pQβ7 or Qβ (+)-RNA was added as template. The yield of the produced Qβ phage particles was 2.4-2.5 × 103 PFU/μL reaction.
|
PFU count 10-2 dilution |
PFU count 10-4 dilution |
PFU count 10-6 dilution |
Yield PFU/μL |
|
|
Positive control Qβ phages |
nd |
nd |
8 |
8 × 105 |
|
CFPS pQβ7 |
~200 |
3 |
0 |
2.5 × 103 |
|
CFPS Qβ (+)-RNA |
~75 |
4 |
0 |
2.4 × 103 |
Table 5: Infectious Qβ virus particles are produced in the homemade CFPS system. A double-layered plaque assay was used to analyse the production of Qβ phage particles in the CFPS systems, which were set up with either pQβ7 DNA (0.8 μg) or Qβ (+)-RNA (24 μg) as the template and incubated at 25°C overnight. Purified Qβ phage particles were included in the plaque assay as a positive control (upper row). “PFU count 10-2 dilution”, “PFU count 10-4 dilution” and “PFU count 10-6 dilution” indicate the observed number of plaque-forming units (PFU) on the plates containing the 10-2, 10-4 and 10-6 dilutions, respectively. The number of plaques was counted, where density allowed a sound determination of the number of PFU. “Yield PFU/μL” provides the number of PFU related to the volume of the CFI systems. “nd”: Not determined due to a high plaque density making the precise counting impossible.
3.4. Analysis of the production of Qβ-encoded proteins in the CFPS system
The Qβ genome encodes the maturation protein (48.5 kDa), the coat protein (14.3 kDa), the read-through protein (36 kDa) and the catalytic RdRP subunit called the β-subunit (65.5 kDa) [8]. The production of viral proteins in the CFPS system was evaluated by including [14C]-serine in the systems to radiolabel the produced proteins. The radioactive samples were analysed on a 4%-20% gradient SDS-gel that was dried and exposed to a storage phosphor screen, which was scanned in a storage phosphor imager. The results show clearly visible bands corresponding to protein sizes of approximately 65 kDa, 14 kDa and 11 kDa as well as a faint band around 45 kDa in the lanes with the CFPS system containing pQβ7 or Qβ (+)-RNA (Figure 3, lanes B and C). The proteins of 14 kDa, 45 kDa and 65 kDa correspond to the molecular weights expected for the coat protein, the maturation protein, and the β-subunit, respectively. The 11 kDa protein, seen in the lanes containing Qβ templates, is an unknown product. No proteins are detected in the lanes containing purified Qβ phage particles (Fig. 3, lane A) or the CFPS system with the control DNA (Fig. 3, lane D), which encodes a 10 kDa protein that cannot be detected using the applied method (Figure 3, lane D).
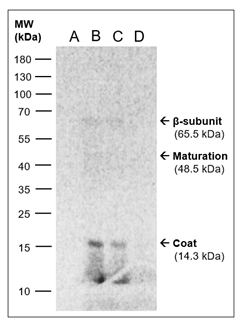
Figure 3: Qβ proteins are produced in the CFPS system. CFPS reactions added [14C]-serine were analysed by SDS-PAGE, followed by phosphorimaging. Contents of the individual lanes: A: Purified Qβ phages (2 × 106 phage particles). B: CFPS reaction with pQβ7 DNA (0.8 μg). C: CFPS reaction with Qβ (+)-RNA (24 μg). D: CFPS reaction with control DNA (0.5 μg). The CFPS systems were incubated at 25°C overnight and 5-μL samples were loaded. Marker: “PageRuler Prestained Protein Ladder” (Thermo Scientific). The arrows indicate the expected positions of the Qβ proteins based on their molecular weights and the molecular weight marker. This figure is cropped, and the full figure is found in Additional file 2.
3.5. Analysis of the replication of Qβ RNA in the CFPS system
The production of Qβ virus particles requires translation of viral proteins, replication of the viral genome and assembly of phage particles [10]. The production of Qβ RNA was analysed by RT-PCR. RNA was extracted from the CFPS reactions added pQβ7 or control DNA as a template. RNA extracted from purified Qβ phage particles was included as a positive control. The purified RNA was DNase treated, and first-strand cDNA synthesis was performed by reverse transcription primed by a reverse primer recognizing the 3’-UTR of the Qβ (+)-RNA. Reactions without RNA template, primers or reverse transcriptase were included as negative controls. The presence of the Qβ genome was analysed in a subsequent PCR reaction using the resulting cDNA as the template. The primers used were a forward primer binding upstream of the β-subunit gene, and a reverse primer binding within the β-subunit gene (Table 1 and Figure 1). The size of the expected PCR product, indicating the presence of Qβ RNA, is 1036 bp. Figure 4 shows that a PCR product of ~1036 bp appears in the reaction based on RNA extracted from the CFPS system with pQβ7 as a template (Figure 4, lane B), while the CFPS reaction with the control DNA does not give rise to this band as expected (Figure 4, lane A). A comparison of lanes A and B also reveals a number of shared bands, which therefore seems to be products generated from RNA species inherent to the CFPS system. The positive control with RNA extracted from purified Qβ phage particles (Fig. 4, lane C) contains the ~1036 bp PCR product, indicating the presence of the Qβ genome, together with numerous nonspecific PCR products. No PCR products are generated in the negative controls lacking either a template (Figure 4, lane D) or reverse transcriptase (Figure 4, lane F), which indicates that the pQβ7 plasmid has been successfully removed from the samples, and the PCR reactions were carried out without contaminations. The control reaction without added primer during first-strand synthesis (Figure 4, lane E) reveal a band at ~1036 Da, which is most likely a result of the ability of the 3’ end of the Qβ (+)-RNA to self-anneal and thereby act as a primer.
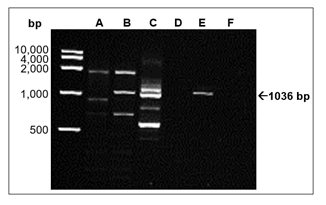
Figure 4: RT-PCR analysis of the production of Qβ RNA in the CFPS system. RNA from the CFPS systems was purified and DNase treated. First-strand synthesis was primed by a primer recognizing the 3’UTR of the Qβ genome (Fig. 1 and Table 1). RNA extracted directly from purified Qβ phages was included as a positive control. Negative controls without primers, template or reverse transcriptase were also included. Next, PCR reactions were run with primers annealing to the Qβ genome, and the products were analysed by 1.5% agarose gel electrophoresis. Contents of the individual lanes: A: CFPS reaction with control DNA. B: CFPS reaction with pQβ7. C: Qβ phages. D: Reverse transcription without added template. E: Reverse transcription without added primer. F: Reverse transcription without reverse transcriptase. Marker: “FastRuler High Range DNA Ladder”. The arrow indicates the size of the expected PCR fragment.
4. Discussion
4.1. Production of infectious phage particles in CFI systems
In the present study, E. coli-based CFI systems were tested for their ability to support Qβ infection. The goal was to develop a model CFI system allowing viral translation and replication, and the production of infectious phage particles to be studied. The commercial protein expression system PURE is based on purified components from E. coli essential for T7 transcription and translation. The potential production of infectious Qβ phage particles in the PURE system was evaluated with a double-layered plaque assay (Table 3). In general, viruses are well-known to exploit a variety of host proteins during infection, and EF-Tu, EF-Ts, S1 and Hfq have been found to be important for replication of Qβ (+)-RNA [18]. Unlike the other mentioned host proteins, Hfq is not directly associated with E. coli transcription or translation, and it was therefore not expected to be included in the PURE system. Thus, PURE reactions were set up with or without added Hfq to evaluate its importance. Hfq was added in 10-1,000 times molar excess relative to added genomes. Surprisingly, the PURE system did not support the production of infectious phage particles irrespective of the presence of Hfq (Table 3). The replication of (+)-RNA viruses is dependent on the interaction between a wide range of host factors with a few viral components during cell entry, gene expression, virus assembly and release. (+)-RNA viruses, especially, rely on the interaction with RNA-binding host proteins [1], and new host proteins employed by viruses are continuously discovered [34]. Thus, it is possible that Qβ is unable to replicate in the PURE system due to the absence of one or more host proteins essential for phage particle production. The host proteins involved in the replication of the Qβ (+)-RNA appear to be well established [35-38], while other steps during the infection cycle (e.g. the assembly of phage particles) may rely on unknown interactions with host proteins. The commercial S30 T7 system was initially tested for its ability to replicate Qβ from an added genome template. The double-layered plaque assay showed that this system successfully produced infectious Qβ phage particles at a yield of 3.5 × 104 to 2.5 × 105 PFU/μL reaction (Tables 3 and 4). The production of Qβ phage particles is solely originating from the added viral genomes, since intact Qβ phage particles were found unable to replicate in the S30 T7 system (Table 4). This observation seems reasonable, since the Qβ (+)-RNA is expected to be inaccessible, when it is encapsulated in the particle. Specifically, the Qβ (+)-RNA is highly structured with stem-loop structures binding to coat-protein dimers and the maturation protein composing the viral capsid [22]. Most likely, the release of Qβ (+)-RNA relies on the interaction between the maturation protein and the F-pilus on the surface of live E. coli cells [39], which is unlikely to occur in a CFI system, where cell walls have been removed by S30 centrifugation.
4.2. Production of viral proteins during Qβ infection in the CFPS system
In addition to the detection of infectious phage particles, the production of viral proteins in the homemade CFPS system was analysed by radiolabelling (Figure 3). The results showed an intense band corresponding to the molecular weight of the coat protein (14.3 kDa) (Figure 3, lane B and C). Additionally, faint bands matching the molecular weights of the β-subunit (65.5 kDa) and the maturation protein (48.5 kDa) were observed (Figure 3, lanes B and C). The ratio of the translated viral proteins is qualitatively similar to results presented in a previous study [10], where the coat protein was produced in approximately 100 times excess relative to the other viral proteins during 45 minutes of infection [10]. In the present study, no attempt was made to quantify the individual viral proteins, but overall, the pattern of translation regulation is comparable to in vivo conditions. The translation is tightly regulated by secondary structures of the viral genome [15], which leaves only the Shine-Dalgarno sequence and the start codon of the coat protein to be accessible for translation. The start codons for the other viral proteins only become accessible during translation of the coat protein or replication of the viral genome [16]. Curiously, an intense band corresponding to a protein of around 11 kDa appeared in the CFPS system with the Qβ genome added (Figure 3, lanes B and C). This protein is not encoded by the Qβ (+)-RNA. Since no transcription inhibitor is added to the reaction, it could potentially be an E. coli protein, such as Hfq, which assists the Qβ RdRP complex during infection. The Hfq monomer has a molecular weight of 11.2 kDa and could potentially be upregulated in E. coli in response to the cell stress caused by the Qβ infection. To this date, there are no studies published on the regulation of Hfq during viral infections. However, Hfq levels have been shown to rise upon increased expression of the Hfq target sRNAs [40], and a similar phenomenon may occur in response to Qβ infection, since Hfq is interacting with the Qβ (+)-RNA [18].
4.3. Detection of Qβ RNA in the CFPS system
The replication of viral RNA in the CFPS system was analysed by RT-PCR using primers specific for the Qβ genome (Figure 4). The expected PCR product of 1036 bp was observed upon analysis of the CFPS reaction with pQβ7 added as a template (Figure 4, lane B), which confirms that the Qβ (+)-RNA has been replicated. In contrast, the CFPS reaction with control DNA as a template did not give rise to the 1036 bp PCR product confirming that this band is a consequence of Qβ template replication (Figure 4, lane A). A comparison of lanes A and B also reveals a number of shared bands, which therefore seems to be products generated from RNA species intrinsic to the system (e.g. rRNA and mRNA). The Qβ (+)-RNA was extracted from purified Qβ phage particles and included in the RT-PCR as a positive control (Fig. 4, lane C). The results revealed the ~1036 bp PCR product, indicating the presence of the Qβ genome, together with several nonspecific PCR products. The appearance of nonspecific PCR products during RT-PCR could be explained by random priming by contaminating cellular nucleic acids [41]. The control reaction without added primer during first-strand synthesis (Figure 4, lane E) resulted in a band at ~1036 Da, that is most likely a result of the ability of the 3’ end of the Qβ (+)-RNA to self-anneal and thereby act as a primer, which is commonly observed for (+)-RNA viruses [42].
4.4. Cell-free infection systems
A few viruses have previously been synthesised in cell-free transcription/translation systems under different conditions and with varying yields. Table 6 shows an overview of some of the previous work together with the results from the CFPS-based CFI system developed in this study.
|
Host extract |
Virus |
Genome |
Genome concentration |
PFU/μL reaction |
Genome replication |
Protein synthesis |
Ref. |
|
E. coli
|
Qβ |
ss (+)-RNA 4,217 nt RNA or DNA copy |
355 nM RNA 2 nM DNA |
2.4 × 103 |
Detected by RT-PCR |
Detected by radio labelling |
This study |
|
E. coli
|
MS2 |
ss (+)-RNA 3,569 nt |
150 nM |
100 |
na |
Detected by radio labelling |
[54]* |
|
E. coli
|
?X174 |
ss (+)-DNA 5,386 nt |
1 nM |
103
|
na |
na |
[48] |
|
E. coli
|
T7 |
ds DNA 39,937 bp |
1 nM |
0.1-5 × 106
|
Detected by agarose gel electro-phoresis |
na |
|
|
E. coli
|
MS2 |
ss (+)-RNA 3,569 nt |
150 nM |
4.23 × 109
|
na |
na |
[44] |
|
E. coli
|
?X174 |
ss (+)-DNA 5,386 nt |
5 nM |
1.9 × 109
|
No replication |
na |
|
|
E. coli
|
T7 |
ds DNA 39,937 bp |
0.25 nM |
3.35 × 108
|
Detected by measurement of PFU with and without dNTPs |
na |
|
|
E. coli
|
T4 |
ds DNA 168,903 bp |
1 nM |
106
|
na |
na |
[55] |
|
HeLa |
Poliovirus |
ss (+)-RNA 7,500 nt |
≈ 3.3 nM (8 ng/μL reaction) |
0.5-45 |
Detected by (-)-strand specific PCR |
Detected by radio labelling |
[45]* |
|
HeLa |
EMCV |
ss (+)-RNA 7,800 nt RNA or DNA copy |
≈ 4 nM RNA (10 ng/μL reaction) ≈ 0.21 mM DNA (0.75 ng/μL reaction) |
1.7 × 107 1.4 × 106
|
Detected by Northern blotting |
Detected by Western blotting |
[56] |
Table 6: Overview of cell-free virus synthesis. “Host extract” lists the host cell from which the extract in the cell-free system is originating. “Virus” indicates the virus, which is produced in the CFI system. “Genome” shows the type and size of the viral genome, while “RNA or DNA copy” indicates that either an RNA or DNA copy has been added as a template in the CFI system, and “Genome concentration” gives the concentration of the added genomic template. ”PFU/μL reaction” indicates the yield of infectious viruses produced per μL cell-free reaction. “Genome replication” and “Protein synthesis” show if viral genome replication and protein synthesis has been analysed. “Ref.” lists the references of the included studies, where “*” indicates the unoptimized CFI systems. This table is not meant to provide a complete overview of all cell-free virus productions. na: Not analysed.
Notably, the present study monitors both replication and translation of the viral genome in the CFI system, in addition to the production of infectious phage particles. The yield of the CFPS system used in this work is 2.4-2.5 × 103 PFU/μL reaction, which is comparable to or higher than the yields obtained in other unoptimised cell-free systems (marked with * in Table 6). It has previously been shown that optimisation of various components of the CFI systems can improve the phage particle yield dramatically. Amongst the possible steps of optimisation is the addition of molecular crowders (e.g. PEG 8000). PEG 8000 mimics the molecular crowding in the cell, which at an optimal concentration can stimulate self-assembly of macromolecular complexes, while an excessive amount of PEG can result in aggregation and misfolding [43]. In the CFPS system presented here, 3% PEG 6000 was included based on Krinsky et al. [31], but an optimisation of the concentration and degree of polymerisation of PEG might increase the yield significantly. Ideally, the concentration of proteins and other essential molecules in the cell extract should be adjusted to mimic the composition of the E. coli cytoplasm [44]. Specifically, an increased concentration of translation factors and an optimisation of NTP concentrations have been shown to increase the yield [45, 46], together with the adjustment of the magnesium ion concentration, since the magnesium concentration affect ribosome functionality [47]. In the present CFPS reaction, the cytoplasmic extract is diluted to contain 24.3 mg/mL protein, compared to ~10 mg/mL protein in other cell-free systems [44, 48], which indicates a potential for further adjustment of concentrations and ratios of individual components.
4.5. Future perspectives
In this study, infectious Qβ phage particles have been produced in a cell-free system. Qβ has previously been used in the development of anti-viral drugs and vaccines [49, 50]. Additionally, Qβ has been used in phage therapy targeting specific pathogenic bacteria, which are eliminated by lysis. This method enables specific targeting of pathogenic bacteria [51]. In the future, CFI systems can potentially be employed in similar biomedical applications. Furthermore, Qβ-based CFI systems may be suitable in evolution studies, since the high mutation rate of the Qβ RdRP complex causes the accumulation of point mutations in the produced RNA genomes [52]. CFI systems are advantageous in allowing the study of interactions between viral and host cell components, which can pose a challenge in vivo e.g. if the host protein of interest is crucial to the cell. Ribosomal protein S1 and Hfq are examples of essential, RNA-binding host proteins that are exploited by the Qβ virus. However, the functional relevance of Hfq and S1 during viral infection cannot be studied by mutagenesis in vivo, since mutations are prone to cause cell death or other unwanted reactions in the cells [19]. This, however, would not be a problem in a cell-free extract depleted of the protein of interest e.g. via degron tagging [53]. Additionally, the in vitro infection systems may facilitate the study of newly discovered bacteriophages [5, 6], and knowledge of their life cycle can be employed for specific medical and biotechnological purposes e.g. in phage therapy, as some of the newly discovered bacteriophages have been isolated as targets of pathogenic bacteria [5, 6]. In general, the expansion of the phage diversity contributes to the study of the impact of viruses on their hosts.
5. Conclusion
Here, we have shown that CFI systems can be established for the bacteriophage Qβ based on a commercial transcription/translation S30 T7 system as well as a custom-made CFPS system [31]. In the latter system, we show that viral proteins are produced with an overall pattern of translation regulation comparable to in vivo conditions. In addition, replication of the viral RNA is demonstrated for this CFI system. Both CFI systems support the self-assembly of infectious Qβ phage particles with yields that are comparable to other unoptimized CFI systems, and the home-made CFI system provides ample opportunity for further improvements via careful titration of crucial components. For future studies, the CFI systems can be used to study known and newly discovered viruses. Understanding the molecular details of viral infections can lead to the development of anti-viral drugs and vaccines, and it can be used in phage therapy.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent to publication
Not applicable.
Availability of data and materials
All data generated or analysed during this study are included in this published article and its supplementary information files.
Competing interests
No competing financial interests exist.
Funding
This research was supported by grants from the Hartmann Foundation, Dagmar Marshall’s Foundation and the Riisfort Foundation, which mainly covered running expenses related to the experimental design and execution.
Author’s contributions
PR planned and conducted all experiments. CK conceived the study and advised on experimental design. Both authors contributed to the writing of the manuscript.
Acknowledgements
We thank Karen Margrethe Nielsen and Lan Bich Van for skilled technical assistance, and associate professor Mikkel Girke Jørgensen (Department of Biochemistry and Molecular biology, University of Southern Denmark) for scientific discussions. The plasmid pTYB11-6 encoding Hfq was kindly provided by associate professor Jakob Møller-Jensen, Department of Biochemistry and Molecular biology, University of Southern Denmark, while purified Hfq was a donation from MSc Manar Merhi, Aarhus University.
Author information
Not relevant.
Affiliations
Department of Molecular Biology & Genetics, University of Aarhus, Gustav Wieds Vej 10 C, DK-8000 Aarhus C, Denmark
References
- Ahlquist P, Noueiry AO, Lee W, et al. Host Factors in Positive-Strand RNA Virus Genome Replication. J Virol 77 (2003): 8181-8186.
- Baltimore D. Expression of Animal Virus Genomes. Bacteriol Rev 35 (1971): 235-241.
- Nagy P, Pogany J. The dependence of viral RNA replication on co-opted host factors. Nat Rev Microbiol 10 (2011): 137-149.
- Pumpens P, Renhofa R, Dishlers A, et al. The True Story and Advantages of RNA Phage Capsids as Nanotools. Intervirology 59 (2016): 74-110.
- Callanan J, Stockdale SR, Shkoporov A, et al. Expansion of known ssRNA phage genomes: From tens to over a thousand. Sci Adv 6 (2020).
- Krishnamurthy SR, Janowski AB, Zhao G, et al. Hyperexpansion of RNA Bacteriophage Diversity. PLOS Biol 14 (2016): 1002409.
- Chetverin AB. Paradoxes of replication of RNA of a bacterial virus. Mol Biol (Mosk) 45 (2011): 160-172.
- Priano C, Arora R, Butke J, et al. A complete plasmid-based complementation system for RNA coliphage Qβ: Three proteins of bacteriophages Qβ (Group III) and SP (Group IV) can be interchanged. J Mol Biol 249 (1995): 283-297.
- Weiner AM, Weber K. Natural read-through at the uga termination signal of Qβ coat protein cistron. Nat New Biol 234(1971): 206-209.
- Tsukada K, Okazaki M, Kita H, et al. Quantitative analysis of the bacteriophage Qβ infection cycle. Biochim Biophys Acta - Gen Subj 1790 (2009): 65-70.
- Kim H, Yin J. Energy-efficient growth of phage Q Beta in Escherichia coli. Biotechnol Bioeng 88 (2004):148-156.
- Meng R, Jiang M, Cui Z, et al. Structural basis for the adsorption of a single-stranded RNA bacteriophage. Nat Commun 10 (2019): 3130.
- Gorzelnik KV, Zhang J. Cryo-EM reveals infection steps of single-stranded RNA bacteriophages. Prog Biophys Mol Biol (2020).
- R?mnieks J, Tars K. Protein-RNA Interactions in the Single-Stranded RNA Bacteriophages. In: Sub-cellular biochemistry (2018): 81-303.
- Weissmann C, Billeter MA, Goodman HM, et al. Structure and Function of Phage RNA. Annu Rev Biochem 42 (1973): 303-328.
- Jacobson AB, Arora R, Zuker M, et al. Structural plasticity in RNA and its role in the regulation of protein translation in coliphage Qβ. J Mol Biol 275 (1998): 589-600.
- Bernardi A, Spahr PF. Nucleotide Sequence at the Binding Site for Coat Protein on RNA of Bacteriophage R17. Proc Natl Acad Sci 69 (1972): 3033-3037.
- Blumenthal T, Carmichael GG. RNA Replication: Function and Structure of Qβ-Replicase. Annu Rev Biochem 48 (1979): 525-548.
- Hajnsdorf E, Boni I V. Multiple activities of RNA-binding proteins S1 and Hfq. Biochimie 94 (2012): 1544-1553.
- Inouye H, Pollack Y, Petre J. Physical and Functional Homology between Ribosomal Protein S1 and Interference Factor i. Eur J Biochem 45 (1974): 109-117.
- Kavita K, de Mets F, Gottesman S. New aspects of RNA-based regulation by Hfq and its partner sRNAs. Curr Opin Microbiol 42 (2018): 53-61.
- Cui Z, Gorzelnik KV, Chang J, et al. Structures of Qβ virions , virus-like particles , and the Qβ – MurA complex reveal internal coat proteins and the mechanism of host lysis 114 (2017): 11697-11702.
- Chakraborty A, Ko C, Henning C, et al. Synchronised infection identifies early rate-limiting steps in the hepatitis B virus life cycle. Cell Microbiol 22 (2020): 13250.
- Vieyres G, Angus AGN, Haberstroh A, et al. Rapid synchronization of hepatitis C virus infection by magnetic adsorption. J Virol Methods 157 (2009): 69-79.
- Al-Fageeh MB, Smales CM. Control and regulation of the cellular responses to cold shock: the responses in yeast and mammalian systems. Biochem J 397 (2006): 247-259.
- Sacha JB, Watkins DI. Synchronous infection of SIV and HIV in vitro for virology, immunology and vaccine-related studies. Nat Protoc 5 (2010): 239-246.
- Kai L, Dötsch V, Kaldenhoff R, et al. Artificial environments for the co-translational stabilization of cell-free expressed proteins. PLoS One 8 (2013): 56637.
- Failmezger J, Rauter M, Nitschel R, et al. Cell-free protein synthesis from non-growing, stressed Escherichia coli. Sci Rep 7 (2017): 16524.
- Shaklee PN, Miglietta JJ, Palmenberg AC, et al. Infectious positive- and negative-strand transcript rnas from bacteriophage Qβ cDNA clones. Virology 163 (1988): 209-213.
- Møller T, Franch T, Højrup P, et al. Hfq: a bacterial Sm-like protein that mediates RNA-RNA interaction. Mol Cell 9 (2002): 23-30.
- Krinsky N, Kaduri M, Shainsky-Roitman J, et al. A simple and rapid method for preparing a cell-free bacterial lysate for protein synthesis. PLoS One 11 (2016): 1-13.
- Cormier J, Janes M. A double layer plaque assay using spread plate technique for enumeration of bacteriophage MS2. J Virol Methods 196 (2014): 86-92.
- Klovins J, Van Duin J. A long-range pseudoknot in Q(β) RNA is essential for replication. J Mol Biol 294 (1999): 875-884.
- Leblanc S, Brunet MA. Modelling of pathogen-host systems using deeper ORF annotations and transcriptomics to inform proteomics analyses. Comput Struct Biotechnol J 18 (2020): 2836-2850.
- Kidmose RT, Vasiliev NN, Chetverin AB, et al. Structure of the Qβ replicase, an RNA-dependent RNA polymerase consisting of viral and host proteins. Proc Natl Acad Sci 107 (2010): 10884-10889.
- Gytz H, Mohr D, Seweryn P, et al. Structural basis for RNA-genome recognition during bacteriophage Qβ replication. Nucleic Acids Res 43 (2015): 10893-108906.
- Takeshita D, Tomita K. Molecular basis for RNA polymerization by Qβ replicase. Nat Struct Mol Biol 19 (2012): 229-237.
- Takeshita D, Yamashita S, Tomita K. Molecular insights into replication initiation by Qβ replicase using ribosomal protein S1. Nucleic Acids Res 42 (2014): 10809-10822.
- Dai X, Li Z, Lai M, et al. In situ structures of the genome and genome-delivery apparatus in a single-stranded RNA virus. Nature 541 (2017): 112-116.
- Vecerek B, Moll I, Bläsi U. Translational autocontrol of the Escherichia coli hfq RNA chaperone gene. RNA 11 (2005): 976-984.
- Craggs JK, Ball JK, Thomson BJ, et al. Development of a strand-specific RT-PCR based assay to detect the replicative form of hepatitis C virus RNA. J Virol Methods 94 (2001): 111-120.
- Tuiskunen A, Leparc-Goffart I, Boubis L, et al. Self-priming of reverse transcriptase impairs strand-specific detection of dengue virus RNA. J Gen Virol 91 (2010): 1019-10127.
- Minton AP. Implications of macromolecular crowding for protein assembly (2000): 34-39.
- Garamella J, Marshall R, Rustad M, et al. The All E. coli TX-TL Toolbox 2.0: A Platform for Cell-Free Synthetic Biology. ACS Synth Biol 5 (2016): 344-355.
- Molla A, Paul AV, Wimmer E. Cell-Free, De Nova Synthesis of Poliovirus. Science (80- ) 254 (1991): 1647-1651.
- Li J, Gu L, Aach J, et al. Improved cell-free RNA and protein synthesis system. PLoS One 9 (2014).
- Laohakunakorn N, Grasemann L, Lavickova B, et al. Bottom-Up Construction of Complex Biomolecular Systems With Cell-Free Synthetic Biology. Front Bioeng Biotechnol 8 (2020): 213.
- Shin J, Jardine P, Noireaux V. Genome replication, synthesis, and assembly of the bacteriophage T7 in a single cell-free reaction. Vol. 1, ACS synthetic biology. United States 1 (2012): 408-213.
- Wang C, Fernández de Ávila BE, Mundaca-Uribe R, et al. Active Delivery of VLPs Promotes Anti-Tumor Activity in a Mouse Ovarian Tumor Model. Small 16 (2020).
- Hong S, Zhang Z, Liu H, et al. B Cells Are the Dominant Antigen-Presenting Cells that Activate Naive CD4(+) T Cells upon Immunization with a Virus-Derived Nanoparticle Antigen. Immunity 49 (2018): 695-708.
- Gordillo Altamirano FL, Barr JJ. Phage Therapy in the Postantibiotic Era. Clin Microbiol Rev 32 (2019).
- Hossain MT, Yokono T, Kashiwagi A. The Single-Stranded RNA Bacteriophage Qβ Adapts Rapidly to High Temperatures: An Evolution Experiment. Viruses 12 (2020): 638.
- Carr AC, Taylor KL, Osborne MS, et al. Rapid depletion of target proteins allows identification of coincident physiological responses. J Bacteriol 194 (2012): 5932-5940.
- Katanaev VL, Spirin AS, Reuss M, et al. Formation of bacteriophage MS2 infectious units in a cell-free translation system. FEBS Lett 397 (1996): 143-148.
- Rustad M, Eastlund A, Jardine P, et al. Cell-free TXTL synthesis of infectious bacteriophage T4 in a single test tube reaction. Synth Biol (Oxford, England) 3 (2018).
- Kobayashi T, Nakamura Y, Mikami S, et al. Synthesis of encephalomyocarditis virus in a cell-free system: from DNA to RNA virus in one tube. Biotechnol Lett 34 (2012): 67-73.
Supplementary File
|
PFU added |
PFU count |
PFU/μL |
|
104 PFU |
nd |
nd |
|
200 PFU |
110 |
5.5 × 105 |
|
100 PFU |
45 |
4.5 × 105 |
|
50 PFU |
32 |
6.4 × 105 |
Table S1: Positive controls included in the double-layered plaque assay. Purified Qβ phages were included in the double-layered plaque assay at the expected PFU: 104, 200, 100 and 50. “PFU count” indicates the observed number of PFU. “PFU/μL” indicates the PFU/μL in the stock of purified Qβ phages. “nd”: Not determined due to a high plaque density making the precise counting impossible.